Researchers at the University of Illinois Urbana-Champaign have announced the development of a transformative neuroimaging and sequencing platform capable of mapping the intricate web of neural connections within the brain at unprecedented speeds. This new methodology, dubbed "Connectome-seq," utilizes molecular barcodes to tag individual neurons, allowing scientists to chart thousands of connections with single-synapse resolution. The breakthrough, led by Boxuan Zhao, a professor of cell and developmental biology, represents a significant leap forward in the field of connectomics—the comprehensive study of the brain’s structural and functional wiring. By converting the physical architecture of the brain into a high-throughput sequencing problem, the team has provided a new lens through which researchers can examine the fundamental circuitry of the central nervous system.
The human brain contains approximately 86 billion neurons, each forming thousands of synaptic connections with other cells. Understanding this vast network is critical for deciphering how the brain processes information, regulates behavior, and succumbs to disease. Traditionally, mapping these connections has been a labor-intensive process, often requiring years of manual labor to reconstruct even small portions of neural tissue. The introduction of Connectome-seq promises to reduce this timeline from years to weeks, offering a scalable solution that could eventually lead to a complete atlas of the mammalian brain. The findings, recently published in the prestigious journal Nature Methods, highlight the technology’s ability to reveal previously unknown cellular links, providing a potential roadmap for therapeutic interventions in conditions ranging from Alzheimer’s disease to psychiatric disorders.
Decoding the Brain’s Architectural Blueprint
At its core, the study of the brain’s circuitry is akin to reverse-engineering a highly sophisticated computer. As Professor Boxuan Zhao noted, understanding the function of a central processing unit (CPU) is impossible without a detailed map of its internal wiring. In the biological context, the "wires" are the axons and dendrites that extend from neurons, and the "junctions" are the synapses where signals are transmitted. Without a comprehensive map of these connections, scientists are limited in their ability to optimize brain function or repair it when neurological disorders occur.
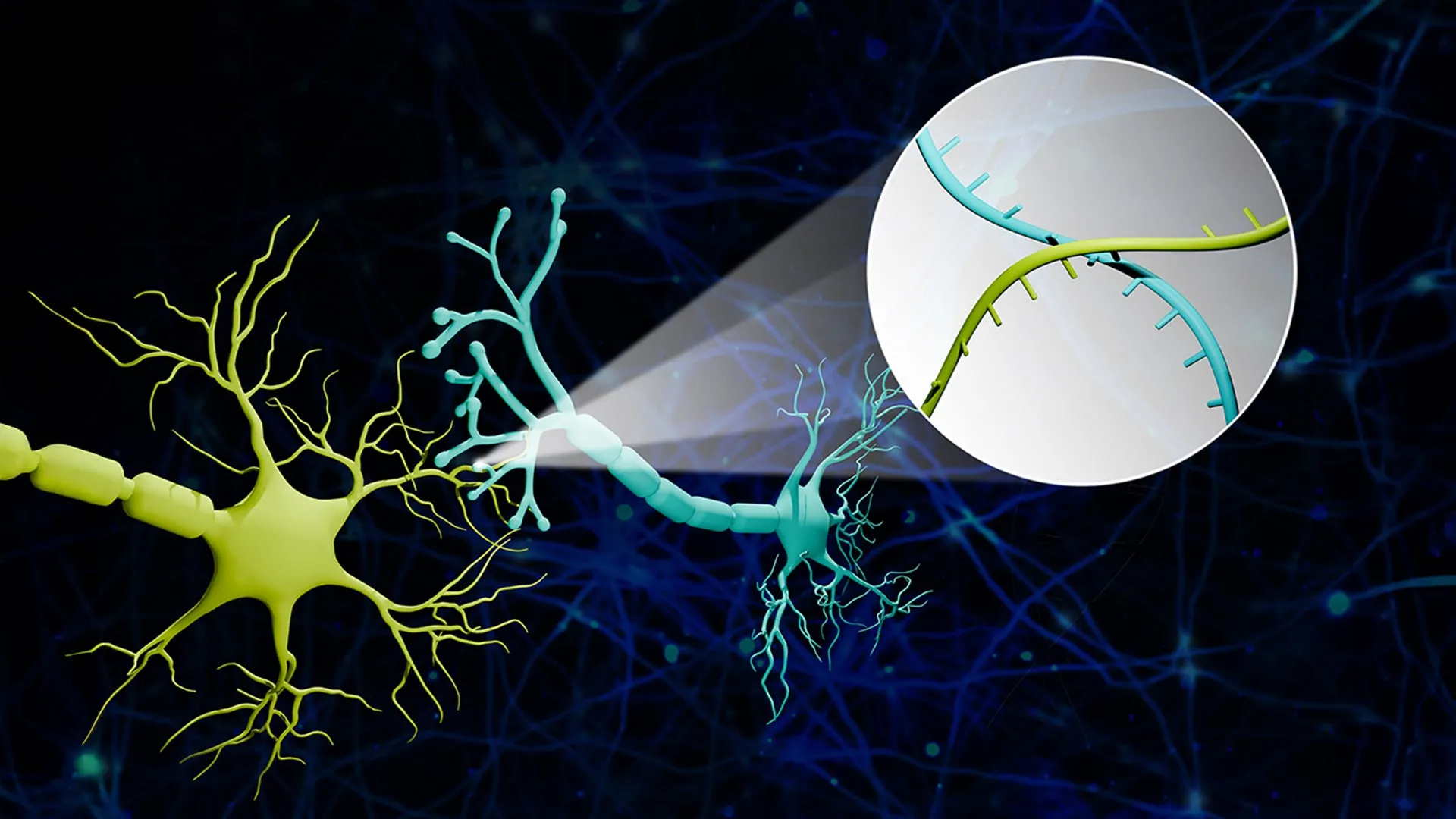
Connectome-seq addresses this challenge by assigning a unique RNA "barcode"—a specific sequence of nucleotides—to every individual neuron. These barcodes serve as digital fingerprints. To ensure these identifiers reach the points of connectivity, the researchers engineered specialized proteins designed to transport the RNA barcodes from the cell body (the soma) down the long axonal projections to the synapse. When two neurons form a connection, their respective barcodes meet at this junction. By isolating these synapses and utilizing high-throughput sequencing technology, the researchers can read the pairs of barcodes found together, effectively identifying which cell was "talking" to which.
The Evolution of Connectomics: From Microtomes to Sequencing
To appreciate the impact of Connectome-seq, one must consider the limitations of traditional brain-mapping techniques. For decades, the gold standard for mapping neural connections has been serial-section electron microscopy (EM). This process involves slicing brain tissue into sections only a few nanometers thick, imaging each slice with an electron microscope, and then using advanced software—or sometimes human volunteers—to manually trace the path of every neuron across thousands of images. While EM provides unmatched detail, it is prohibitively slow and expensive. Mapping a single cubic millimeter of mouse brain tissue using these methods can generate petabytes of data and take years of computational and manual effort.
In recent years, sequencing-based tools have emerged as an alternative. These tools can label many neurons simultaneously, but they have historically struggled with resolution. Many existing sequencing methods can show where a neuron’s axon extends generally within the brain, but they cannot confirm with certainty whether a functional synapse has been formed with a specific neighbor. Connectome-seq bridges this gap by focusing specifically on the synapse. By isolating the physical meeting points of neurons, the technology ensures that the data collected reflects actual connectivity rather than mere proximity.
Unveiling the Pontocerebellar Circuit: A Case Study in Discovery
To demonstrate the efficacy of the platform, Zhao’s team applied Connectome-seq to the pontocerebellar circuit in a mouse brain. This circuit is a vital pathway that connects the pons—a part of the brainstem involved in relaying signals—to the cerebellum, which is responsible for motor control, coordination, and some cognitive functions. The team successfully mapped over 1,000 neurons within this circuit, achieving a level of detail that would have been unattainable with previous sequencing technologies in such a short timeframe.
The results of this initial mapping were surprising even to the researchers. The analysis revealed connectivity patterns that had never been documented in the adult mouse brain. Specifically, the team identified direct links between certain cell types that were previously thought to be independent or only indirectly connected. These findings suggest that the brain’s wiring is even more complex than current anatomical models suggest. The discovery of these "hidden" connections demonstrates the power of Connectome-seq to refine our understanding of neural architecture and highlights the limitations of older, less granular mapping techniques.
Strategic Implications for Alzheimer’s and Neurodegenerative Disease
The primary clinical value of Connectome-seq lies in its potential to transform our approach to neurodegenerative diseases. Conditions like Alzheimer’s, Parkinson’s, and Amyotrophic Lateral Sclerosis (ALS) are increasingly viewed as "disconnection syndromes," where the primary pathology involves the progressive breakdown of synaptic links long before the actual death of the neurons themselves.
By comparing the connectomes of healthy brains with those of individuals in various stages of neurodegeneration, researchers can identify the "weak links" in the circuit. Professor Zhao emphasized that catching these changes early is the key to effective treatment. If scientists can identify the specific synapses that are most vulnerable at the onset of Alzheimer’s, they may be able to develop targeted therapies to strengthen those connections. This "circuit-guided" approach could potentially slow or even halt the progression of the disease before cognitive symptoms, such as memory loss or confusion, become irreversible.
Furthermore, because Connectome-seq is fast and relatively inexpensive compared to electron microscopy, it allows for large-scale comparative studies. Researchers can analyze hundreds of different brain samples to see how connectivity varies based on genetics, age, or environmental factors. This statistical power is essential for understanding psychiatric conditions like schizophrenia or autism, which are thought to arise from subtle "miswiring" during brain development rather than the gross anatomical damage seen in late-stage dementia.
The Intersection of Bioinformatics and Sequencing
The success of Connectome-seq relies heavily on the translation of biological structures into digital data. Professor Zhao uses a "balloon analogy" to explain this complex process to the public. If a thousand balloons are floating in a room, each with unique stickers on their strings, and those strings are tied together in various knots, the task of mapping them manually would be overwhelming. Connectome-seq essentially "snips the knots" and reads the stickers. If a single knot contains stickers from Balloon A and Balloon B, it is an irrefutable sign that those two balloons were connected.
This transition from a physical problem to a sequencing problem allows neuroscience to tap into the rapid advancements of the genomics industry. As the cost of DNA and RNA sequencing continues to drop and the speed of data processing increases, the capabilities of Connectome-seq will naturally improve. The team is already working on enhancing the resolution and throughput of the platform, with the ultimate goal of mapping the entire mouse brain—a feat that would involve billions of connections.
Chronology of Development and Future Horizons
The development of Connectome-seq was a multi-year effort involving interdisciplinary collaboration between cell biologists, engineers, and bioinformaticians. The project began with the engineering of the RNA-binding proteins required to transport the barcodes. This was followed by rigorous testing in cell cultures to ensure that the barcodes did not interfere with normal cellular function. Once the molecular machinery was perfected, the team moved to in vivo testing in mouse models, specifically targeting the pontocerebellar circuit to validate the technology against known anatomical benchmarks.
Looking forward, the researchers at the University of Illinois Urbana-Champaign are confident that the platform will become a standard tool in neuroscience laboratories worldwide. The scalability of the technology means it could be used not just for mice, but eventually for larger mammalian brains, providing the first high-resolution maps of the primate connectome. Such data would be invaluable for understanding the unique evolutionary traits that define the human brain.
Institutional Support and Collaborative Frameworks
The research was made possible through significant financial and institutional support. A primary driver of the project was a Neuro-omics Initiative grant from the Wu Tsai Neurosciences Institute at Stanford University. Additional funding was provided by the Elsa U. Pardee Foundation and the Edward Mallinckrodt Jr. Foundation. These organizations focus on high-risk, high-reward research that has the potential to break new ground in the treatment of cancer and neurological diseases.
The collaborative nature of this work underscores a growing trend in science: the "Neuro-omics" movement. This field seeks to integrate the tools of genomics, proteomics, and transcriptomics with traditional neuroscience to create a holistic view of brain function. Connectome-seq is a flagship example of this integration, showing that the future of brain research lies at the intersection of multiple scientific disciplines.
As the scientific community begins to adopt Connectome-seq, the focus will likely shift toward creating open-source databases of brain maps. By sharing these "circuit diagrams," researchers around the globe can collaborate on identifying the structural foundations of intelligence, memory, and emotion. In the long term, this technology may not only help us fix the brain when it breaks but also provide the fundamental knowledge needed to enhance human cognition and quality of life.







Leave a Reply