The pharmaceutical industry grapples with an intractable challenge: the escalating cost and protracted timelines of drug development, coupled with persistently high clinical failure rates. Bringing a new drug from initial discovery to market approval typically consumes a decade to a decade and a half, with some projects extending even longer. This arduous journey is not only time-consuming but exorbitantly expensive, often ranging from hundreds of millions to several billions of dollars per successful drug. Compounding this issue is the phenomenon dubbed "Eroom’s Law" (Moore’s Law spelled backward), which starkly illustrates that the inflation-adjusted cost of drug development has roughly doubled every nine years, indicating a severe decline in R&D efficiency despite technological advancements.
The Pervasive Efficacy Gap: A Core Challenge
A primary driver of these clinical setbacks is the fundamental issue of inadequate efficacy. A comprehensive analysis of clinical trial data spanning from 2010 to 2017 revealed that a staggering 40% to 50% of drugs that fail in clinical trials do so because they simply do not work as intended. This statistic underscores a critical disconnect: the preclinical assays and models currently employed often fail to accurately reflect a drug’s true biological impact and efficacy in complex physiological systems, particularly human patients. The sophisticated biological interactions and disease pathology present in a living organism are notoriously difficult to replicate in simplified laboratory settings.
AI’s Promise and Persistent Hurdles
In recent years, the field of artificial intelligence (AI) has emerged as a beacon of hope for revolutionizing drug discovery. Its potential to accelerate target identification, optimize molecule design, and predict drug-target interactions has attracted substantial investment and widespread optimism. According to a sector review by DealForma, the AI drug discovery sector attracted an impressive 612 venture rounds and approximately $19.9 billion in total capital during 2024 and 2025 alone. Despite this significant influx of resources and intellectual capital, AI has yet to demonstrably move the needle on the industry’s entrenched 90% clinical failure rate. While AI has undoubtedly streamlined certain aspects of early-stage research, its impact on predicting in vivo efficacy and reducing late-stage attrition remains largely unvalidated and mixed. This reality points to deeper, systemic issues within preclinical validation that even advanced computational power has struggled to overcome.
The "Snapshot Assay Problem" and Its Limitations
At the heart of many clinical trial failures lies a fundamental limitation of conventional drug discovery assays: their inherently static nature. Current methodologies frequently reduce the intricate, evolving dynamics of cellular behavior into discrete, single-point measurements or "snapshots." These assays often provide a glimpse of a cellular state at a specific moment in time, failing to capture the continuous, time-dependent responses that characterize real biological processes.
For instance, techniques like transcriptome profiling, while powerful for understanding gene expression, typically involve the destruction of cells. This destructive nature precludes the ability to track the dynamic changes in gene expression within an individual cell over multiple time points. Such methods offer an average picture across a population of cells at a given instant, but they obscure the subtle, heterogeneous, and sequential biological events that dictate a drug’s true mechanism of action and its ultimate efficacy. This lack of dynamic insight means that crucial early-stage responses or delayed effects, which might signal a drug’s therapeutic potential or impending toxicity, are often missed.
The inherent limitations of target-centric drug discovery, which focuses on modulating a single molecular target, have led to a resurgence of interest in phenotypic drug discovery. This approach shifts the focus from individual targets to observing complex cellular behaviors and disease-relevant phenotypes. By screening compounds based on their ability to induce a desired change in cell morphology, growth, or function, phenotypic approaches aim to identify hits that exert a beneficial effect, even if the precise molecular target is initially unknown. However, phenotypic screens present their own set of challenges. These include the difficulty of validating the "hits" identified, the laborious process of "target deconvolution" (identifying the specific molecular targets responsible for the observed phenotype), and the general struggle to translate complex phenotypic signals into clear, actionable mechanistic insights. Without understanding how a drug achieves its phenotypic effect, optimizing it and predicting off-target effects becomes significantly more challenging.
Introducing Live Cell Dynamics (LCD): A Paradigm Shift
Against this backdrop of persistent challenges, a novel approach termed Live Cell Dynamics (LCD) is emerging as a potential game-changer. Developed by scientists at Soley Therapeutics, LCD is a self-supervised machine learning pipeline designed to extract rich, dose- and time-dependent cellular state information directly from continuous brightfield images. Critically, this method operates without the need for traditional stains or labels, which can themselves perturb cellular biology or be difficult to apply in high-throughput, long-term studies. The innovative methodology behind LCD was detailed in a January 2026 paper published in Scientific Reports.
Kurosh Ameri, co-founder and CSO of Soley Therapeutics, elucidated the core philosophy behind LCD: "By treating cellular response as time-resolved information rather than a static snapshot, LCD enables mechanism classification, compound comparison, and detection of complex biology through measurable trajectories." He emphasized the transformative potential of this shift, explaining, "This provides early forward-looking biological signal rather than a late binary readout, shifting drug discovery from observing damage to forecasting a drug’s direction and future impact." This approach fundamentally redefines how researchers observe and interpret drug-cell interactions, moving beyond mere endpoint assessment to a dynamic, predictive understanding.
Robust Validation and Key Findings
The Scientific Reports study provided compelling evidence for LCD’s capabilities. Researchers pre-trained the LCD model on a diverse library of 189 compounds and subsequently evaluated its performance on 81 additional "held-out" compounds, encompassing 10 distinct mechanisms of action. For this initial validation, a single, well-characterized human osteosarcoma cell line (U2OS) was utilized under controlled laboratory conditions.
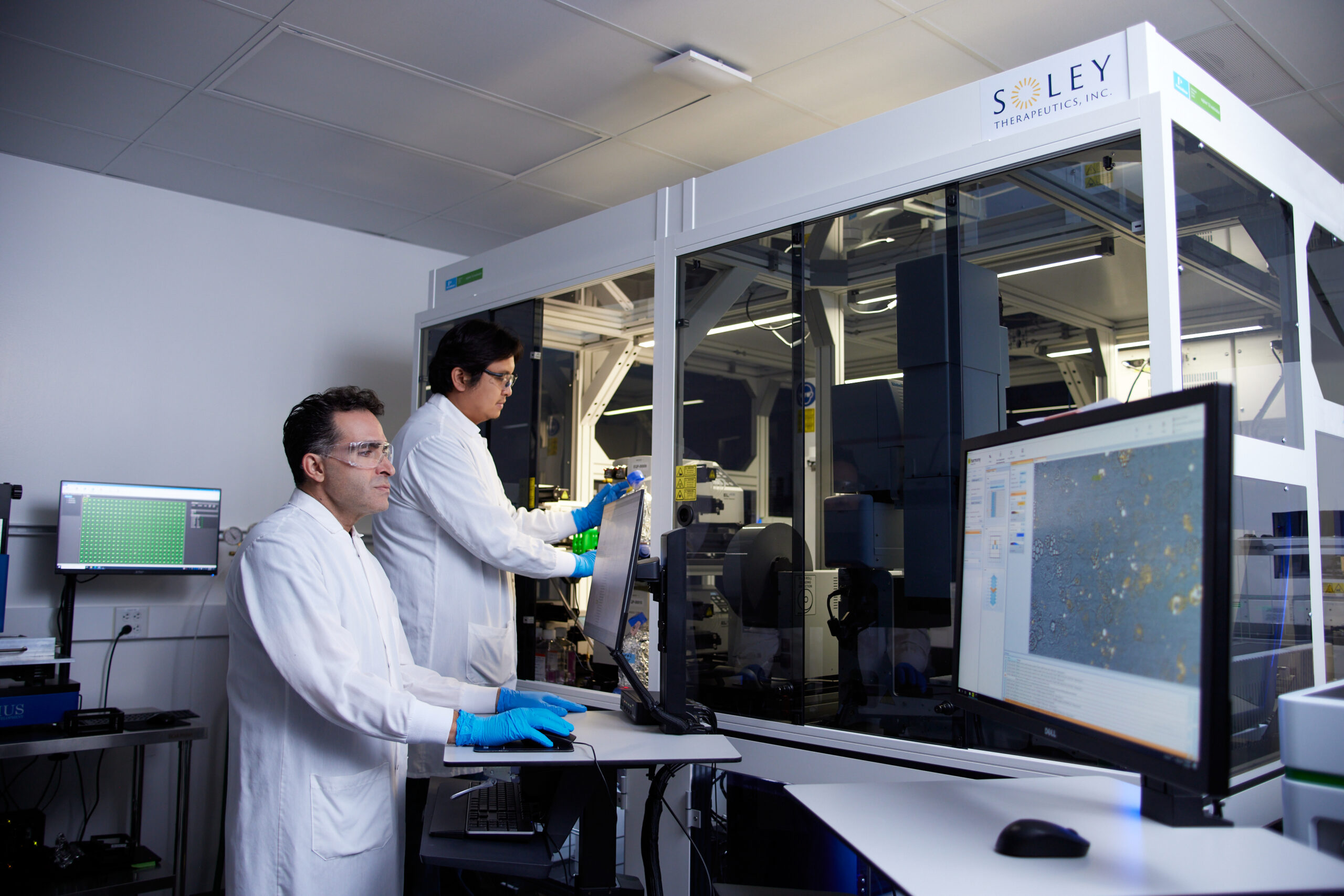

The results were striking: LCD consistently outperformed conventional methods, including simple cell counting and feature extraction based on CellProfiler software, in detecting phenotypic activity across a range of doses and time points. The advantages of LCD were particularly pronounced at early time points and low doses, precisely when traditional methods often struggle to detect subtle biological shifts. This ability to capture nascent cellular responses is critical, as it can provide much earlier indications of a compound’s potential efficacy or toxicity, allowing for quicker decision-making in the drug discovery pipeline.
Furthermore, the study demonstrated that incorporating multiple doses and time points incrementally enhanced LCD’s ability to classify mechanisms of action. This multi-dimensional data allowed LCD to disentangle mechanisms that might appear similar or converge in their effects at later stages of cellular response, but exhibit distinct temporal profiles at earlier points. Ameri reiterated this advantage, stating, "Learned representations from LCD preserved signal in those early regimes and performed strongly across dose and time, while the CellProfiler baseline tended to be comparable only later, or lower at early time points." This highlights LCD’s capacity to extract richer, more nuanced biological information that is otherwise obscured by static or endpoint analyses.
Unmasking Polypharmacology: A Hidden Challenge
One of the most significant and notoriously difficult challenges in drug discovery is the detection of polypharmacology—the phenomenon where a single drug affects multiple biological targets simultaneously. While often considered a hurdle, polypharmacology can also be therapeutically beneficial, particularly in complex diseases like cancer where multi-target intervention is desired. Conventionally, identifying polypharmacology necessitates extensive, costly, and time-consuming assay panels, often involving a broad range of biochemical and cell-based tests.
Remarkably, the LCD model, using only brightfield imaging data, demonstrated the ability to flag both Aurora kinase and JAK inhibitor activity. This detection was entirely consistent with prior studies that required extensive kinome profiling—a highly specialized and resource-intensive technique to assess drug interactions across a vast array of kinases—to reach the same conclusions. This capability of LCD to infer complex polypharmacological profiles from simple, label-free images represents a potentially transformative advancement, offering a far more efficient and cost-effective way to characterize drug-target interactions comprehensively.
Overcoming the Challenges of Brightfield Imaging
Brightfield microscopy, while non-invasive and label-free, presents its own set of technical challenges for quantitative analysis. As Ameri noted, "Brightfield is difficult because the signal is subtle, not evident to the naked eye, contrast is low, and small changes in optics, focus, plate position, or day-to-day setup can create batch effects that swamp biology." These technical variations can easily mask genuine biological signals, making robust data extraction a formidable task.
To overcome these inherent difficulties, the Soley Therapeutics paper outlined two crucial training innovations integrated into the LCD pipeline:
- Plane-agnostic augmentation: This technique trains the model to recognize and prioritize genuine biological features over artifacts related to the focal plane. By introducing variations in focus during training, the model learns to identify consistent cellular changes regardless of slight shifts in image acquisition, thereby making it more robust to experimental variability.
- Cross-batch sampling: This innovation forces the model to learn features that remain stable and consistent across different experimental runs and batches. By exposing the model to data from various experimental setups, it develops the ability to distinguish true biological signals from technical noise or batch-specific effects, significantly improving the reliability and reproducibility of the extracted information.
These technical advancements are critical for ensuring that LCD can reliably operate in the high-throughput, diverse experimental environments characteristic of modern drug discovery. The ability to extract meaningful, stable biological insights from subtle brightfield images without interference from technical noise is a testament to the sophistication of the machine learning architecture.
Future Outlook and Broader Implications
The study’s results collectively demonstrate that "LCD can represent compound behavior as a profile across dose and time, not a single label. Those profiles contain enough structure to separate closely related mechanisms and expose mixed activity, which is exactly the kind of complexity that shows up in development," Ameri concluded. This shift from simplistic, single-point readouts to comprehensive, dynamic profiles marks a significant step towards understanding drug action in a more biologically relevant manner.
However, the current study, while highly promising, utilized a single, well-characterized cancer cell line (U2OS) under tightly controlled laboratory conditions. This specificity means that LCD’s performance and generalizability in more complex, biologically diverse, and disease-relevant models remain to be fully explored. The central question that this foundational work leaves open is whether the observed performance advantages in a controlled compound library will hold up across the inherently messier and more heterogeneous biology found in primary cells, patient-derived organoids, or other advanced disease models. The transition from a simplified in vitro system to models that more closely mimic human disease is often where promising preclinical technologies encounter their greatest challenges.
According to Soley Therapeutics, the immediate next steps involve expanding the application of LCD to a broader range of cell types, including primary human cells and disease-relevant models that incorporate greater biological complexity. Furthermore, the company aims to broaden the coverage of mechanisms of action that LCD can effectively characterize and to integrate the technology for prospective use in active drug discovery programs. Only after LCD has been rigorously validated in settings that more closely approximate human disease can definitive claims about its potential clinical impact and its ability to significantly reduce the industry’s efficacy problem be fairly evaluated.
Should LCD prove successful in these more complex biological contexts, its implications for drug discovery are profound. By providing earlier, more accurate, and dynamically rich biological signals, LCD could significantly accelerate lead optimization, reduce the reliance on costly and time-consuming animal models, and ultimately decrease the high attrition rates in clinical trials. This could translate into a substantial reduction in the cost and time required to bring new therapies to patients, offering a much-needed boost to innovation in an industry burdened by Eroom’s Law. Moreover, its ability to detect polypharmacology and differentiate subtle mechanistic variations could facilitate the development of more precise and effective treatments, particularly for complex diseases where single-target approaches have often fallen short. The journey for Live Cell Dynamics has just begun, but its potential to transform how we discover and develop life-saving medicines is undeniably compelling.











Leave a Reply