New research emerging from Arizona State University (ASU) has unveiled two remarkable and previously unrecognized strategies by which bacteria can move and spread across surfaces, even when their primary modes of locomotion are compromised. These groundbreaking discoveries, detailed in separate studies published in leading scientific journals, challenge long-held assumptions about bacterial mobility and have significant implications for understanding and combating infectious diseases, as well as for harnessing beneficial microbial activities.
Sugar-Fueled "Swashing": A Metabolic Current of Movement
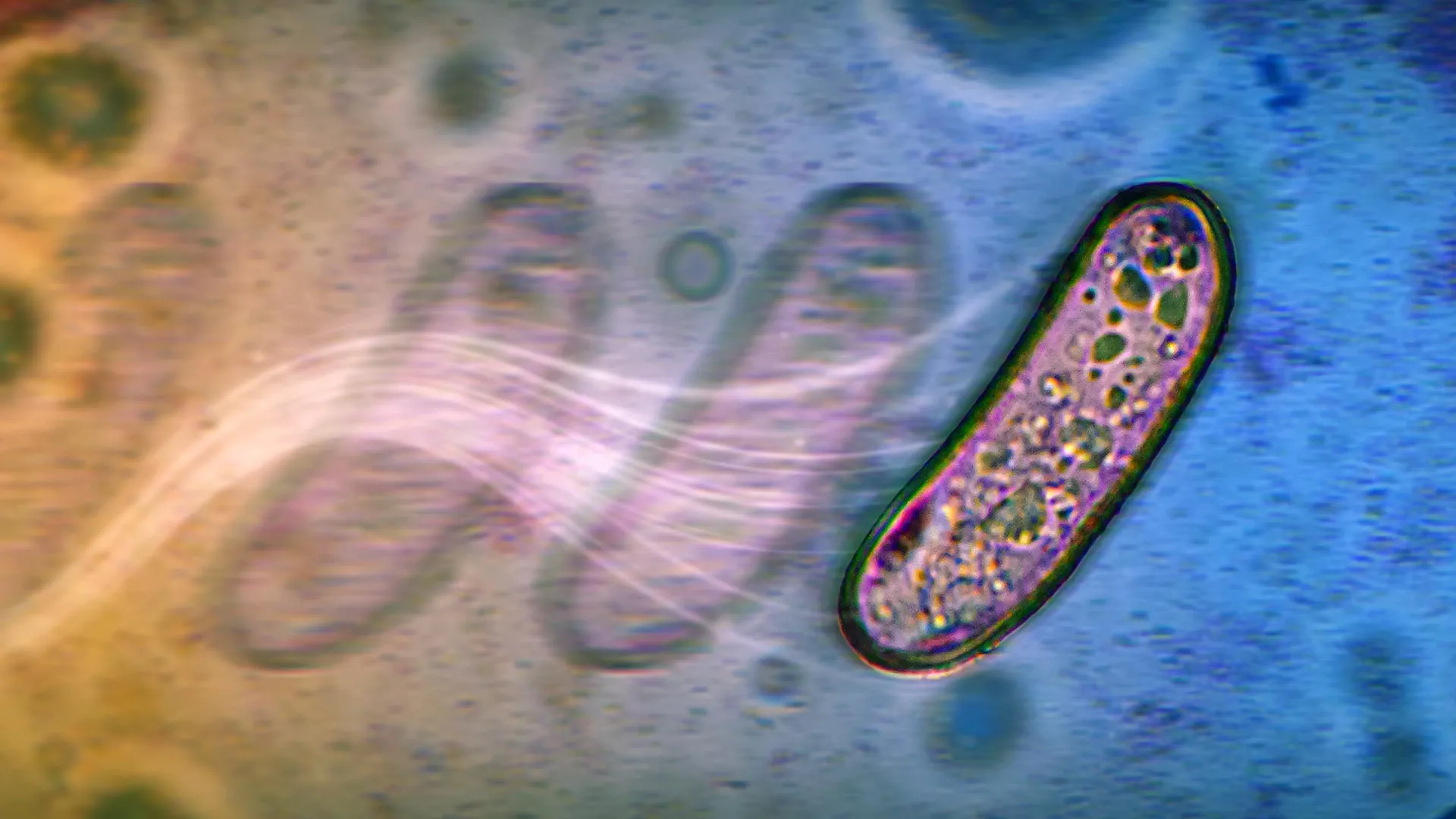
The first study, led by researcher Navish Wadhwa from ASU’s Biodesign Center for Mechanisms of Evolution and Department of Physics, reveals a novel form of bacterial migration dubbed "swashing." This phenomenon allows bacteria like Salmonella and E. coli to spread across moist surfaces without the functional assistance of their flagella, the whip-like appendages typically responsible for propulsion. The research, published in the Journal of Bacteriology and recognized as an Editor’s Pick, indicates that bacterial metabolism itself can generate the forces necessary for movement.
"We were amazed by the ability of these bacteria to migrate across surfaces without functional flagella," stated Wadhwa. "In fact, our collaborators originally designed this experiment as a ‘negative control,’ meaning that we expected, once rendered flagella-less, the cells to not move. But the bacteria migrated with abandon, as if nothing were amiss, setting us off on a multiyear quest to understand how they were doing it."
The swashing mechanism hinges on the fermentation of sugars. When bacteria metabolize sugars such as glucose, maltose, or xylose, they produce acidic byproducts like acetate and formate. These compounds, in turn, influence the surrounding liquid on the surface. They create localized osmotic gradients that draw water towards the bacterial colony, generating subtle outward-flowing currents. These currents, akin to a miniature river system, then passively propel the bacterial cells and their expanding colonies across the substrate.
This metabolic propulsion system was initially an unexpected finding. The experimental setup was designed to isolate the role of flagella by creating flagella-less mutants of Salmonella and E. coli. The expectation was that these modified bacteria would remain stationary, serving as a baseline for comparison. However, to the researchers’ surprise, the flagella-deficient strains exhibited significant movement, prompting a deeper investigation into the underlying mechanisms. The multiyear quest to decipher this unexpected motility eventually led to the identification of the swashing phenomenon.
The implications of swashing are far-reaching, particularly in the context of healthcare and food safety. Disease-causing microbes could potentially colonize medical devices, such as catheters and implants, as well as wounds and food processing equipment, through this metabolic propulsion, even if their flagella are somehow inhibited or absent. This discovery suggests that current strategies focused solely on disabling flagella may not be sufficient to prevent bacterial spread in all scenarios.
"It just goes to show that even when we think we’ve got something figured out, there are often surprises waiting just under the surface, or in this case, above it," Wadhwa added, underscoring the dynamic and often unpredictable nature of microbial life.
The study found that swashing is critically dependent on the availability of fermentable sugars. Without these energy sources, bacteria cannot generate the necessary fluid flows. This highlights the importance of the microenvironment in facilitating bacterial dissemination. For instance, sugar-rich environments within the human body, such as mucus linings in the gut or respiratory tract, could provide ideal conditions for swashing and subsequent infections.
Furthermore, the researchers observed that the addition of surfactants, molecules that reduce surface tension and are found in detergents, effectively halted swashing. Interestingly, these same surfactants did not impede "swarming," another well-documented flagella-driven bacterial motility that enables rapid surface spread. This distinction strongly suggests that swashing and swarming operate via fundamentally different physical principles. The ability to differentiate between these modes of movement could pave the way for targeted interventions. If swashing is the primary mode of spread, environmental modifications or specific chemical agents that disrupt metabolic byproduct generation or fluid dynamics might be more effective than flagella-targeting treatments.
The findings have direct relevance for sanitation protocols in the food industry. Recognizing that bacteria like E. coli and Salmonella, notorious for causing foodborne illnesses, can spread through passive fluid flows could lead to enhanced cleaning and disinfection strategies in food processing facilities. Manipulating environmental factors such as surface pH or sugar concentrations could become a key strategy to curb bacterial proliferation and spread. Initial experiments indicated that even minor adjustments to acidity could significantly influence bacterial movement via swashing.
Beyond food safety, similar moist environments within the human body—including gut mucus, wound exudates, and the urinary tract—present potential niches for swashing-mediated colonization and infection, even in the absence of functional flagella. This broadens the scope of potential applications for this research in clinical settings.
Molecular Gearbox: Navigating Surfaces with Precision
The second ASU study, conducted by researchers from the Biodesign Center for Fundamental and Applied Microbiomics, the Biodesign Center for Mechanisms of Evolution, and the School of Life Sciences, delves into a different mode of bacterial locomotion in a group of bacteria known as flavobacteria. These microbes do not possess flagella for swimming. Instead, they rely on a sophisticated molecular machine called the type 9 secretion system (T9SS) to move along surfaces. This system functions as a molecular conveyor belt that propels the cell forward.
The T9SS enables flavobacteria to glide by moving an adhesive-coated belt around the cell’s exterior. This process effectively "pulls" the bacterium along the surface, a motion compared to a microscopic snowmobile. The research, published in the journal mBio and highlighted by its authors as a significant breakthrough, has now elucidated a critical regulatory component of this system.
Specifically, the team identified a protein within the T9SS, known as GldJ, which acts as a molecular gear shifter. This protein controls the direction of the motor that drives the conveyor belt. By making a minor modification to GldJ, researchers found that the motor’s rotation could be reversed from counterclockwise to clockwise. This alteration directly impacts the bacterium’s direction of travel, demonstrating a remarkable level of control over its motility.
"We are very excited to have discovered an extraordinary dual-role nanogear system that integrates a feedback mechanism, revealing a controllable biological snowmobile and showing how bacteria precisely tune motility and secretion in dynamic environments," explained lead researcher Dr. Anjelika Shrivastava. "Building on this breakthrough, we now aim to determine high-resolution structures of this remarkable molecular conveyor to visualize, at atomic precision, how its moving parts interlock, transmit force and respond to mechanical feedback."
This intricate molecular gearbox allows flavobacteria to adapt their movement in response to complex and changing environments, offering an evolutionary advantage by enabling more effective navigation. This precise control over motility is crucial for survival and colonization in diverse niches.
The T9SS system’s influence extends beyond just bacterial movement. It plays a multifaceted role in host-microbe interactions, with both detrimental and beneficial consequences depending on the specific microbial community. In the oral microbiome, T9SS-containing bacteria have been implicated in gum disease. The proteins they secrete can trigger inflammatory responses in the mouth, and research has suggested potential links to systemic health conditions like heart disease and Alzheimer’s.
Conversely, in the gut microbiome, T9SS activity can be advantageous. Proteins released through this system can shield antibodies from degradation, thereby bolstering immune defenses and potentially enhancing the efficacy of oral vaccines. This dual role underscores the complexity of microbial systems and their intricate interactions with their hosts.
Understanding the mechanics of this molecular gearbox offers promising avenues for therapeutic development. Researchers envision ways to disrupt the T9SS to prevent bacteria from forming biofilms—sticky, resilient communities that are a major cause of persistent infections and contamination of medical devices. Simultaneously, these insights could be leveraged to support beneficial microbes and design targeted microbiome therapies.
"Unraveling this intricate design will not only deepen our understanding of microbial evolution but also inspire the development of next-generation bioengineered nanomachines and therapeutic technologies," Dr. Shrivastava added, highlighting the potential for this fundamental research to drive innovation in medicine and biotechnology.
Converging Discoveries: A Multifaceted Approach to Bacterial Control
While the "swashing" phenomenon and the T9SS "molecular gearbox" appear to be distinct mechanisms operating on different scales, both discoveries underscore a critical overarching theme: bacteria possess a remarkable repertoire of adaptable strategies for movement and survival. The more diverse the methods available to microbes for spreading, the more challenging it becomes to control them.
These findings collectively suggest that traditional approaches to combating bacterial infections, which often focus on disabling flagella, may need to be re-evaluated and broadened. The existence of alternative motility mechanisms means that simply blocking flagella could be insufficient to halt the spread of pathogens.
The research points toward the critical importance of controlling the environmental conditions in which bacteria reside. Factors such as the availability of fermentable sugars, the precise pH levels of surfaces, and the chemical composition of the surrounding environment could all play significant roles in modulating bacterial movement. Furthermore, interfering with specific molecular systems like the T9SS gearbox could offer a dual benefit: preventing bacterial motility and inhibiting the secretion of harmful proteins that contribute to disease.
As scientists continue to unravel the complex world of bacterial locomotion and interaction, these ASU studies represent significant strides in understanding these microscopic architects of life and their profound impact on our world. The insights gained promise to inform new strategies for disease prevention, enhance public health measures, and unlock novel biotechnological applications.











Leave a Reply