Evolution, the relentless engine of biological innovation, has long been nature’s most sophisticated engineering process. Within the microscopic world of cells, a constant flux of DNA, RNA, and protein variations arises, with natural selection acting as a discerning architect, favoring those organisms that exhibit superior functionality and adaptability. Humanity’s engagement with this fundamental process dates back millennia, with early agricultural practices serving as a rudimentary yet profoundly impactful form of directed evolution. By selectively breeding crops and livestock, our ancestors consciously guided the evolutionary trajectory of these species, promoting the proliferation of traits that enhanced productivity and resilience.
Today, this ancient principle has been refined and amplified in the laboratory through a powerful technique known as directed evolution. Scientists leverage this methodology to meticulously improve the performance of critical biological molecules, most notably proteins such as enzymes and antibodies. These engineered proteins are indispensable across a vast spectrum of applications, from life-saving pharmaceuticals and intricate industrial manufacturing processes to the formulation of everyday consumer products like laundry detergents.
The Limitations of Traditional Directed Evolution
Despite its remarkable successes, conventional directed evolution methodologies are not without their inherent constraints. A primary limitation lies in their typical reliance on a constant selection pressure. This approach inherently favors proteins that maintain a consistently high level of activity across all conditions. However, this idealized scenario rarely mirrors the complex reality of biological systems. Many proteins within living cells function not as static powerhouses but as dynamic signaling molecules, molecular switches, or intricate "logic gates." These proteins are designed to respond to fluctuating environmental cues, integrating multiple inputs to make binary, yes-or-no decisions. Consequently, their efficacy hinges on their ability to fluidly transition between distinct states.
Consider, for instance, a protein that must transiently activate, then deactivate, and subsequently re-activate under specific circumstances. When evolutionary experiments are solely designed to reward a single, persistent state of activation, other essential functional states can degrade. This can lead to a critical loss of a protein’s capacity to switch appropriately, a deficiency that can have severe detrimental consequences for cellular viability. In extreme cases, such dysregulation can lead to cell death. The inherent difficulty in engineering proteins with complex, multi-state dynamic behavior has therefore remained a significant hurdle for existing directed evolution approaches.
A Novel Light-Based Strategy for Protein Evolution
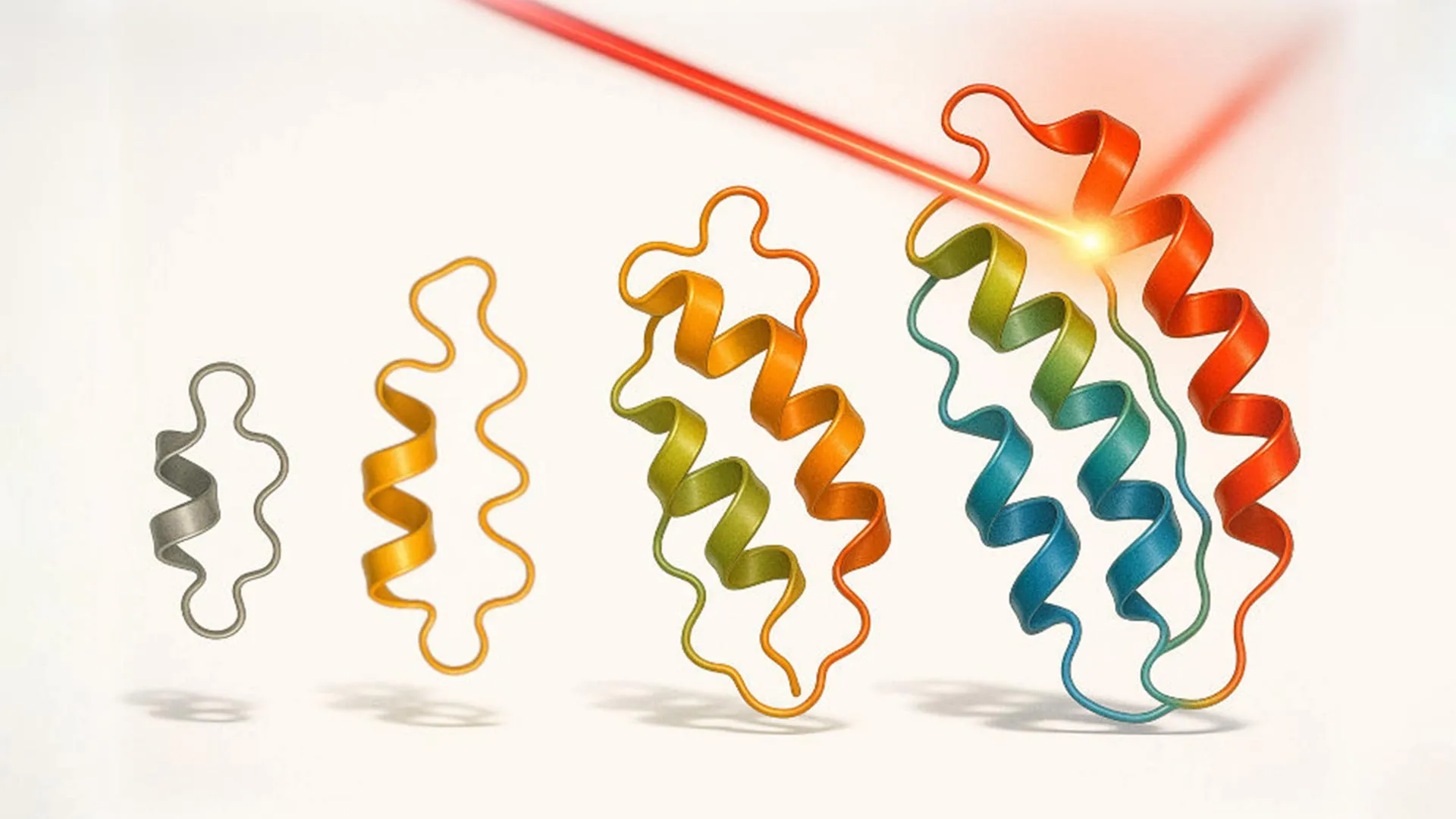
In a groundbreaking development, researchers led by Sahand Jamal Rahi at EPFL’s Laboratory of the Physics of Biological Systems have unveiled an innovative new approach termed "optovolution." This sophisticated method harnesses the precision of light to guide the evolutionary refinement of proteins. The objective is to engineer proteins capable of performing dynamic functions and even executing elementary computational tasks that adhere to binary, yes-or-no logic.
The seminal study, published in the esteemed journal Cell, represents a significant stride towards aligning directed evolution with the nuanced operational principles of natural cellular systems. Within living organisms, the precise timing of biological events and the seamless switching between functional states are often as crucial as the intrinsic strength of a molecular signal.
Engineering Yeast Cells to Select Optimal Protein Variants
To construct their pioneering system, the research team employed the versatile budding yeast, Saccharomyces cerevisiae. This organism, a staple in both the brewing industry and the scientific research community, provided an ideal model system. The scientists ingeniously re-engineered the yeast cell cycle, establishing a dependency where cell division was contingent upon the dynamic behavior of the protein undergoing evolution. Specifically, the protein was required to transition cleanly between active and inactive states for the cell to successfully propagate.
The researchers ingeniously coupled the protein’s output signal to a critical regulator governing the cell cycle. This regulator plays an indispensable role during a specific phase of the cell cycle but becomes profoundly toxic during another. Consequently, if the evolved protein remained locked in either an "on" or "off" state for an extended duration, the yeast cell would either stall its division process or perish. Only those yeast cells harboring proteins that demonstrated the ability to switch states at the precisely correct temporal junctures were able to continue dividing, thereby ensuring their survival and reproduction.
Utilizing Light for Real-Time Control of Evolution
Light emerged as the key enabler for precise control over this intricate evolutionary process. The research team leveraged optogenetics, a cutting-edge technique that allows for the precise activation or deactivation of genes through the application of light. By delivering precisely timed pulses of light, the scientists were able to compel the protein of interest to alternate between its distinct functional states.
Each cycle of the yeast cell division process, which typically spans approximately 90 minutes, served as a rapid, high-throughput assessment. This "pass or fail" test meticulously evaluated whether the protein had successfully switched states at the opportune moment. Proteins that exhibited superior dynamic switching capabilities facilitated the survival and reproduction of their host cells, while variants demonstrating suboptimal switching were systematically eliminated. This automated selection mechanism, powered by optovolution, enabled the identification and enrichment of proteins with enhanced dynamic behavior without the need for laborious manual screening or iterative adjustments by the researchers.
Emergence of Novel Protein Variants and Expanded Color Sensitivity
Through the application of optovolution, the research team successfully evolved several distinct classes of proteins. Their initial focus was on refining a widely utilized light-controlled transcription factor. The team generated an impressive 19 novel variants of this transcription factor. These new variants exhibited several desirable improvements, including enhanced sensitivity to light, significantly reduced activity in the absence of light (darkness), and, notably, the ability to respond to green light, a departure from the typically blue-light dependent systems. The engineering of proteins that can respond to warmer colors of light, such as green or red, has historically been considered an exceptionally challenging endeavor. This difficulty stems from the inherent light absorption properties of the protein machinery involved, which often favors shorter, bluer wavelengths.
In a further demonstration of the technique’s versatility, the scientists also engineered a red light optogenetic system. This advancement eliminated the requirement for yeast cells to be supplemented with an external chemical cofactor. The evolutionary process serendipitously resulted in a mutation that effectively disabled a native yeast transport protein. This unexpected evolutionary outcome allowed the optogenetic system to efficiently utilize light-sensitive molecules that were already endogenously present within the cell, thereby simplifying the experimental setup and enhancing the ease of use for researchers.
Proteins Engineered to Function as Miniature Computers
Beyond light-sensing proteins, the study conclusively demonstrated that optovolution’s capabilities extend to engineering proteins that can perform more complex computational functions. The researchers successfully evolved a transcription factor that operates akin to a single-protein computational unit. This engineered protein was designed to activate gene expression only when two distinct input signals were simultaneously present: one mediated by a light signal and the other by a chemical signal. This represents a significant step towards creating sophisticated cellular circuits capable of processing information.
The ability of proteins to exhibit dynamic behavior is fundamental to a vast array of biological processes. These include sensing subtle changes in the environment, making critical decisions within the intricate network of cellular signaling pathways, and orchestrating the precise control of cell division. By enabling these dynamic behaviors to evolve organically within living cells, optovolution opens up unprecedented avenues for advancements in synthetic biology, biotechnology, and fundamental biological research.
Broader Implications and Future Directions
The implications of this research are far-reaching. The optovolution technique holds immense potential for assisting scientists in the design of more intelligent and sophisticated cellular circuits. It could pave the way for the development of novel optogenetic tools that can respond independently to different colors of light, offering finer control and greater specificity in experimental manipulations. Furthermore, this approach provides a powerful new lens through which to understand the evolutionary mechanisms that give rise to complex protein behaviors.
The ability to engineer proteins with dynamic, multi-state functionality and computational capabilities is crucial for a variety of future applications. In medicine, such proteins could lead to more targeted drug delivery systems that respond to specific physiological cues, or to diagnostic tools capable of detecting disease markers with unprecedented sensitivity. In industrial biotechnology, engineered enzymes with enhanced dynamic regulation could lead to more efficient and sustainable manufacturing processes.
The research team’s findings suggest a paradigm shift in how we approach protein engineering. Moving beyond static optimization, the focus now shifts to the dynamic interplay of protein function and cellular context. This evolution in methodology is likely to accelerate discoveries across multiple scientific disciplines, offering solutions to long-standing challenges and unlocking new possibilities for the manipulation and understanding of biological systems. The ability to mimic nature’s elegant solutions for dynamic control and computation within engineered proteins marks a significant milestone in our quest to harness the power of biology.













Leave a Reply