The pharmaceutical industry faces an escalating crisis marked by soaring costs, protracted timelines, and persistently low success rates in bringing new medicines to patients. Traditional drug discovery journeys typically span 10 to 15 years, sometimes even longer, from initial discovery to regulatory approval. This lengthy process is compounded by astronomical financial burdens, with the cost of developing a single drug ranging from hundreds of millions to multiple billions of dollars. Alarmingly, the inflation-adjusted cost of drug development has roughly doubled every nine years, a phenomenon dubbed "Eroom’s Law" – the inverse of Moore’s Law – illustrating a disturbing trend of diminishing returns in pharmaceutical innovation.
A primary driver of this inefficiency and significant clinical trial failures is a pervasive lack of efficacy. A comprehensive analysis of clinical trial data from 2010 to 2017 revealed that between 40% and 50% of drugs that fail in clinical trials do so because they simply do not work as intended. This stark statistic underscores a fundamental disconnect: preclinical assays, the cornerstone of early-stage drug development, often poorly reflect a drug’s true biological activity and therapeutic potential in living systems. The consequence is a staggering 90% clinical failure rate that has remained largely unimproved despite considerable efforts and technological advancements.
The Unyielding Challenges of Modern Drug Discovery
The pharmaceutical sector’s productivity dilemma, encapsulated by Eroom’s Law, is a multifaceted problem. Beyond the sheer financial and temporal investment, the complexity of human biology and disease pathology presents immense hurdles. Many diseases, particularly chronic and complex ones like cancer, neurodegenerative disorders, and autoimmune conditions, involve intricate biological pathways and redundant compensatory mechanisms that are difficult to model accurately in simplified laboratory settings. This inherent biological complexity often leads to unforeseen side effects or, more commonly, a lack of desired therapeutic effect when a drug moves from controlled preclinical environments to heterogeneous human populations.
Moreover, the regulatory landscape has become increasingly stringent, demanding higher standards of safety and efficacy. While essential for patient protection, these requirements add further layers of time and cost to the development process. The economic pressure is immense, pushing pharmaceutical companies to seek revolutionary approaches that can de-risk their pipelines and improve the predictability of success.
AI’s Promise and Current Reality in Drug Discovery
In recent years, the field of artificial intelligence (AI) has emerged as a beacon of hope for transforming drug discovery. Attracting substantial investment and widespread optimism, the AI drug discovery sector recorded 612 venture rounds and approximately $19.9 billion in total capital between 2024 and 2025, according to DealForma’s sector review. Proponents argue that AI can accelerate target identification, optimize compound design, predict molecular interactions, and even aid in clinical trial design, thereby dramatically shortening timelines and reducing costs.
Despite this influx of capital and enthusiastic adoption, AI’s demonstrable impact on the abysmal 90% clinical failure rate remains limited and mixed. While AI has shown promise in specific areas like virtual screening and lead optimization, translating these early-stage computational successes into tangible improvements in clinical efficacy and approval rates has proven challenging. This gap highlights that even advanced computational power often relies on — and is therefore limited by — the quality and relevance of the biological data it processes. If the underlying experimental data fails to capture the intricate, dynamic nature of cellular responses, even the most sophisticated AI algorithms may struggle to make accurate predictions about a drug’s performance in a complex biological system.
The Limitations of Conventional Assays: A "Snapshot" Problem
A fundamental impediment to improving drug discovery success lies in the structural limitations of conventional drug discovery assays. For decades, the dominant paradigm has been to reduce complex, evolving cellular behavior into static, single-point measurements – what experts now refer to as the "snapshot assay problem." These methods often capture a cell’s state at a single time point, typically after a fixed period of drug exposure, providing an incomplete and often misleading picture of the drug’s interaction with its biological target.
For instance, common assays for gene expression profiling, such as transcriptome analysis, require the destruction of cells. This destructive nature makes it exceedingly difficult, if not impossible, to track the dynamic gene expression patterns of an individual cell over multiple time points, thereby obscuring the temporal nuances of drug response. Similarly, many phenotypic assays, while designed to observe complex cellular behaviors, still often rely on end-point measurements or sparse sampling, failing to capture the full trajectory of cellular changes induced by a therapeutic agent.
These limitations have spurred a resurgence of interest in phenotypic drug discovery (PDD), an approach that focuses on observing observable changes in cellular behavior rather than modulating a single, predefined molecular target. PDD gained prominence in the early days of pharmacology but receded with the rise of target-centric drug discovery, which sought to identify and modulate specific molecular targets implicated in disease. However, the disappointing returns from target-based approaches have led researchers back to PDD. While PDD offers the advantage of identifying novel mechanisms of action and drug candidates that might be missed by target-based screens, it introduces its own set of challenges. These include difficulties in hit validation, the complex process of target deconvolution (identifying the specific molecular targets responsible for the observed phenotype), and translating broad phenotypic signals into precise mechanistic insights. Without a clear understanding of how a drug elicits its effect, further optimization and development become significantly harder.
Introducing Live Cell Dynamics (LCD): A Paradigm Shift
Addressing these critical gaps, Soley Therapeutics has introduced a groundbreaking methodology called Live Cell Dynamics (LCD). Published in a January 2026 Scientific Reports paper, LCD represents a significant leap forward by leveraging a self-supervised machine learning pipeline to extract rich, dose- and time-dependent cellular state information directly from continuous brightfield images. Crucially, this method operates without the need for cumbersome and potentially perturbing stains or labels, preserving the native biological environment of the cells.
Kurosh Ameri, co-founder and CSO of Soley Therapeutics, elucidates the core innovation: “By treating cellular response as time-resolved information rather than a static snapshot, LCD enables mechanism classification, compound comparison, and detection of complex biology through measurable trajectories.” This approach marks a fundamental shift in perspective. Instead of merely observing the end-state damage or a fixed response, LCD allows researchers to forecast a drug’s direction and future impact by analyzing the dynamic path of cellular changes. This provides an early, forward-looking biological signal, offering unprecedented insight into how a drug interacts with living cells over time.
The implications of this shift are profound. By observing cellular trajectories, LCD can potentially identify subtle, early-stage responses that predict later efficacy or toxicity, enabling faster decision-making and de-risking of drug candidates much earlier in the discovery process. This moves beyond a binary readout of "active" or "inactive" to a nuanced understanding of the kinetics and nature of cellular perturbation.
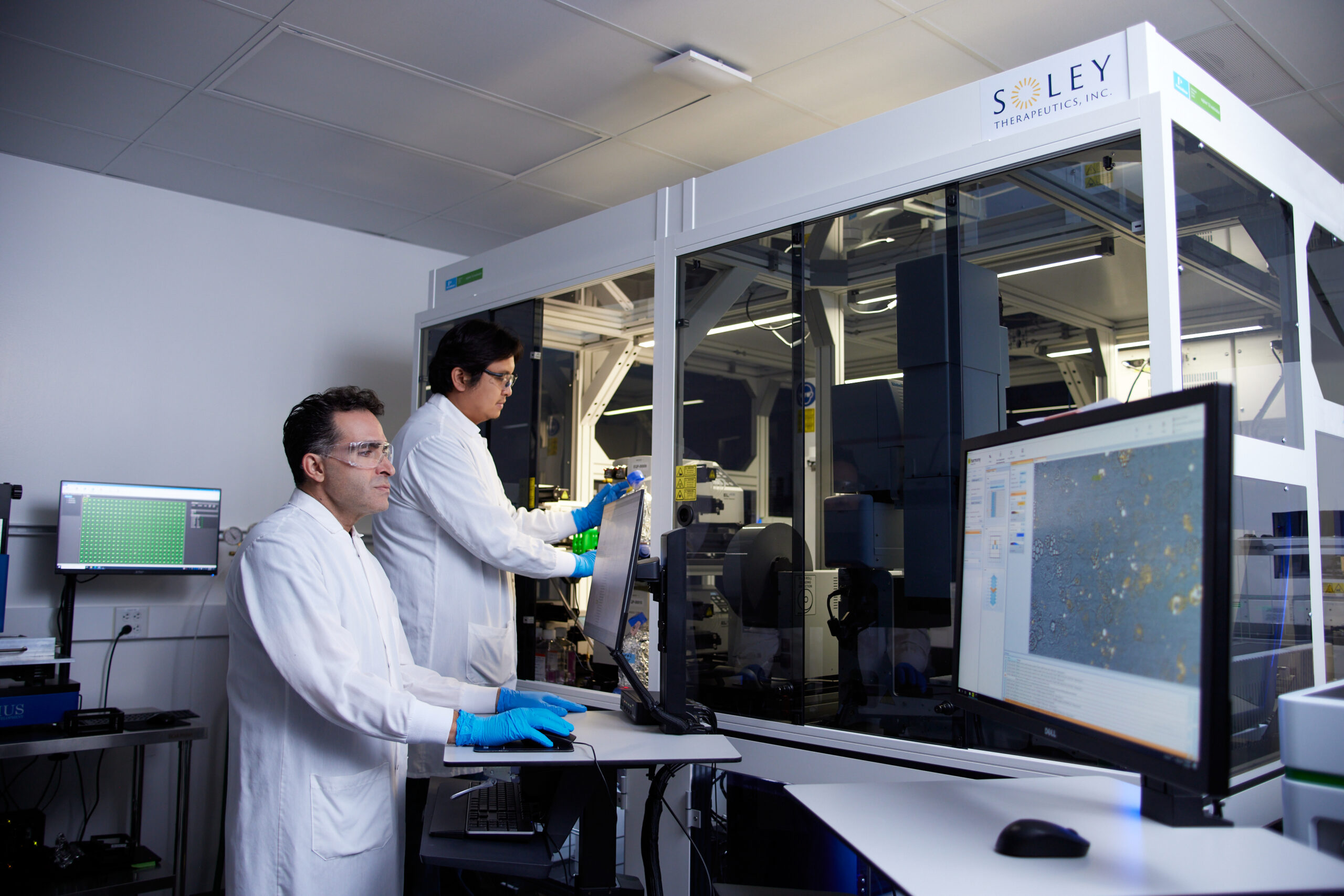

Soley Therapeutics’ Pioneering Research and Key Findings
The study detailed in Scientific Reports rigorously evaluated LCD’s capabilities. Researchers pre-trained the model on a diverse library of 189 compounds and subsequently assessed its performance on an additional 81 held-out compounds, spanning 10 distinct mechanisms of action. All experiments were conducted using a single, well-characterized human osteosarcoma cell line (U2OS), providing a controlled environment for initial validation.
The results were compelling. LCD demonstrably outperformed traditional cell count and CellProfiler-based feature extraction methods for phenotypic activity detection across all doses and time points tested. The advantages were particularly pronounced at early time points and lower doses, where conventional methods often fail to capture subtle biological signals. This early detection capability is critical, as it allows researchers to identify promising compounds or potential issues before significant cellular changes or damage occur. Furthermore, the study revealed that incorporating multiple doses and time points incrementally improved mechanism-of-action classification, allowing LCD to disentangle mechanisms that might appear similar or converge at later stages of cellular response. This ability to resolve closely related mechanisms is a significant advantage for drug developers aiming for highly specific therapeutic interventions.
Ameri emphasized this point, stating, "Learned representations from LCD preserved signal in those early regimes and performed strongly across dose and time, while the CellProfiler baseline tended to be comparable only later, or lower at early time points." This highlights LCD’s superior sensitivity and predictive power, particularly in the crucial early phases of drug-cell interaction.
Beyond basic activity detection, LCD exhibited remarkable capabilities in identifying polypharmacology – the phenomenon where many drugs affect multiple biological targets simultaneously. Polypharmacology is common in therapeutics, contributing to both desired broader effects and undesirable off-target side effects, but it is notoriously difficult to detect using conventional methods, often requiring extensive and costly assay panels. Using only brightfield imaging, the LCD model accurately flagged both Aurora kinase and JAK inhibitor activity, consistent with prior studies that had necessitated extensive kinome profiling to reach the same conclusions. This ability to uncover complex, multi-target interactions from simple, label-free images represents a substantial efficiency gain and could help avoid costly late-stage failures due to unexpected off-target effects.
Ameri acknowledged the inherent technical challenges of brightfield imaging: "Brightfield is difficult because the signal is subtle, not evident to the naked eye, contrast is low, and small changes in optics, focus, plate position, or day-to-day setup can create batch effects that swamp biology." To overcome these hurdles, the paper outlined two ingenious training innovations:
- Plane-agnostic augmentation: This technique trains the model to recognize underlying biological changes irrespective of minor variations in focal plane, preventing it from mistaking optical artifacts for biological signals.
- Cross-batch sampling: This method forces the model to learn features that are stable and reproducible across different experimental runs, effectively separating genuine biological signals from technical noise and day-to-day variations in experimental setup.
These innovations are critical to making LCD a robust and reliable tool for high-throughput drug screening. The results underscore that "LCD can represent compound behavior as a profile across dose and time, not a single label. Those profiles contain enough structure to separate closely related mechanisms and expose mixed activity, which is exactly the kind of complexity that shows up in development,” Ameri concluded. This comprehensive profiling capability offers a far richer and more predictive understanding of a compound’s behavior than static, single-point measurements.
Expert Perspectives and Broader Implications
The emergence of LCD offers a compelling vision for overcoming some of the most entrenched challenges in drug discovery. By providing dynamic, label-free insights into cellular responses, LCD could significantly reduce attrition rates in preclinical development. Identifying non-efficacious or problematic compounds earlier saves immense time and resources, allowing researchers to focus on more promising candidates. This shift from reactive observation to proactive forecasting aligns perfectly with the industry’s desperate need for more predictive models.
The ability of LCD to detect subtle changes at low doses and early time points could revolutionize lead optimization. Instead of relying on brute-force chemical modifications and iterative testing, scientists could use LCD to quickly assess how small structural changes influence the kinetics and trajectory of cellular response, leading to more rational drug design. Furthermore, LCD’s capacity to identify polypharmacology from simple brightfield images could open new avenues for drug repurposing and the development of multi-target therapies, which are increasingly relevant for complex diseases.
Beyond specific drug candidates, LCD has broader implications for understanding fundamental cell biology. By continuously monitoring cellular dynamics in response to various perturbations, researchers can gain deeper insights into disease mechanisms, cellular resilience, and adaptive responses. This could inform the development of more targeted and personalized medicine approaches, where treatments are tailored to an individual’s unique cellular response profile. For instance, by profiling patient-derived cells, LCD might help predict individual responses to therapies, guiding treatment selection and optimizing dosages.
Future Directions and Remaining Hurdles
While the initial findings from Soley Therapeutics are highly promising, the study was conducted under controlled laboratory conditions using a single, well-characterized cancer cell line (U2OS). This means LCD’s performance in more complex, heterogeneous, and clinically relevant models remains to be fully elucidated. The central question the work leaves open is whether the observed performance advantages in a controlled compound library will hold up across the "messier, more heterogeneous biology" of primary cells, patient-derived organoids, or intricate disease models.
According to Soley Therapeutics, the immediate next steps involve expanding LCD to additional cell types, including primary cells and disease-relevant models, to validate its versatility and robustness. This includes broadening the mechanism coverage and integrating LCD into active drug programs for prospective use. Such validation is crucial before any definitive claims can be made about LCD’s clinical impact. The journey from a promising laboratory technique to a widely adopted clinical tool requires rigorous testing in settings that closely mimic human disease conditions. This includes demonstrating its utility in identifying biomarkers, stratifying patient populations, and predicting therapeutic outcomes in diverse biological contexts.
Conclusion
The persistent efficacy problem in drug discovery, compounded by Eroom’s Law and the limitations of static assays, demands innovative solutions. Live Cell Dynamics (LCD) offers a compelling new paradigm by harnessing the power of self-supervised machine learning to extract rich, dynamic biological information from label-free live-cell imaging. The pioneering work by Soley Therapeutics demonstrates LCD’s superior ability to detect early biological signals, classify mechanisms of action, and even uncover complex polypharmacology, offering a more predictive and mechanistic understanding of drug-cell interactions. While the technology’s full clinical impact awaits validation in more complex disease models, LCD represents a significant stride towards de-risking the drug discovery pipeline, accelerating innovation, and ultimately, delivering more effective medicines to patients in need. The shift from static snapshots to dynamic trajectories could indeed be the key to fixing drug discovery’s efficacy problem, heralding a new era of intelligent, data-driven pharmaceutical development.








Leave a Reply