A pioneering team of researchers from ETH Zurich and Karolinska Institutet has unveiled a groundbreaking cross-linking MALDI mass spectrometry (MS) workflow, marking a significant stride in modern drug discovery. This innovative method is engineered to simultaneously capture both the functional response and target binding of a drug candidate within a single assay, directly addressing a critical "information gap" that has long plagued pharmaceutical research and development. The development promises to streamline the drug discovery pipeline, reduce the high failure rates of experimental compounds, and ultimately accelerate the delivery of effective new medicines to patients.
The Persistent Challenge in Drug Discovery: The Information Gap
For decades, the arduous journey of drug discovery has been characterized by a fundamental schism in how potential therapeutic agents are evaluated. Researchers traditionally rely on two distinct, yet incomplete, categories of assays: functional assays and binding assays. While indispensable, each offers only a partial view of a drug candidate’s true potential, often leading to ambiguous results and costly dead ends.
The Dichotomy of Traditional Screening
Functional assays are designed to determine whether a compound elicits a desired biological effect, such as inhibiting an enzyme or activating a receptor. They answer the crucial question of "does it work?" However, they often fall short in explaining how it works, providing little insight into the specific molecular target or mechanism of action. Conversely, binding assays focus on whether a drug candidate physically interacts with its intended biological target, addressing "does it bind?" Yet, a strong binding affinity does not automatically translate into a beneficial functional outcome. A compound might bind to a target effectively but fail to produce the desired therapeutic effect, or worse, trigger unintended side effects due to off-target interactions.
This inherent dichotomy creates an "information gap" that has profound implications for the success rate of drug candidates. The inability to fully understand the interplay between binding and function from early screening stages is a major contributing factor to the alarmingly high attrition rates in clinical development.
High Failure Rates and Economic Burden
The pharmaceutical industry invests billions of dollars and many years into bringing a single new drug to market, a process fraught with uncertainty. Clinical efficacy, or the lack thereof, accounts for a staggering 40% to 50% of clinical failures. This means that nearly half of all drug candidates that enter human trials fail because they simply do not work as intended in patients, despite promising preclinical data. Such late-stage failures are immensely costly, both in terms of financial investment—with estimates for developing a new drug ranging from $1 billion to $2.5 billion—and in lost time, which can extend over a decade or more. Identifying ineffective compounds earlier in the discovery process could save vast resources, allowing companies to reallocate efforts towards more promising avenues.
The Enigma of Protein-Protein Interactions (PPIs)
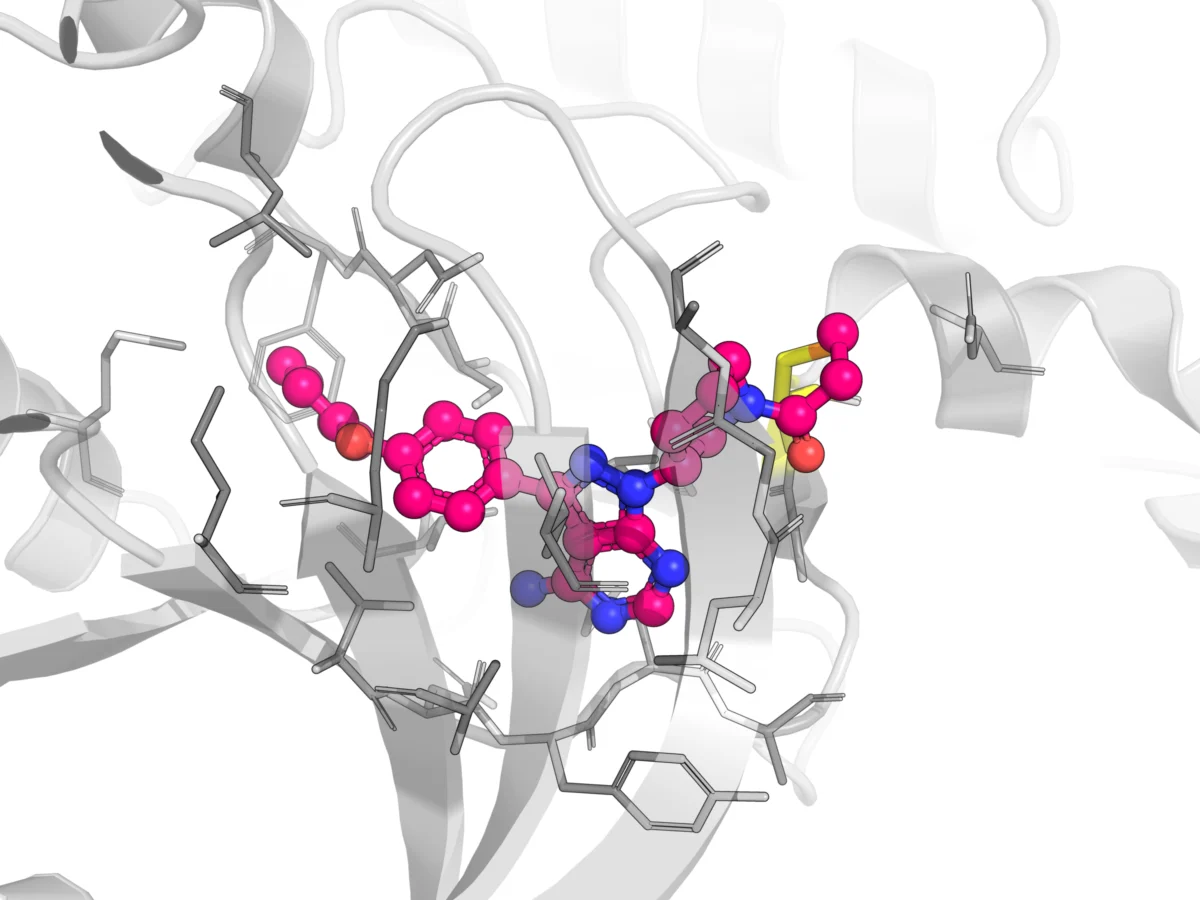
The challenge is particularly acute when dealing with protein-protein interactions (PPIs), which represent a vast and largely untapped class of therapeutic targets. PPIs are fundamental to nearly every biological process, including cell signaling, immune response, and disease progression. Modulating dysregulated PPIs holds immense potential for treating a wide array of diseases, from cancer to neurodegenerative disorders.
However, targeting PPIs with small-molecule drugs has proven exceptionally difficult. Unlike enzyme active sites or receptor binding pockets, PPI interfaces are typically large, flat, and often transient, lacking the well-defined "druggable" pockets that conventional drug discovery approaches rely upon. This inherent complexity makes it challenging to design molecules that can specifically and effectively disrupt or stabilize these interactions, further exacerbating the information gap and contributing to the high failure rate of PPI-modulating drug candidates. The new cross-linking MALDI-MS workflow directly confronts this challenge by offering a novel way to analyze these intricate interactions.
A Novel Approach: Cross-Linking MALDI Mass Spectrometry
The innovation from the ETH Zurich and Karolinska Institutet team lies in a clever modification of an established analytical technique: Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry (MALDI-MS). By introducing a crucial chemical cross-linking step, the researchers have transformed MALDI-MS from a primary identification and measurement tool into a powerful integrated drug screening platform.
Reimagining MALDI-MS
Conventional MALDI-MS is a widely used analytical technique in biochemistry and molecular biology. It excels at measuring the masses of macromolecules like proteins and peptides, making it invaluable for applications such as protein identification, enzyme activity assays, and quality control in protein production. The technique works by embedding the analyte in a crystalline matrix, which is then irradiated with a laser. The matrix absorbs the laser energy, transferring it to the analyte, causing its ionization and desorption into the gas phase, where ions are then separated and detected based on their mass-to-charge ratio.
However, a significant limitation of conventional MALDI-MS, particularly in the context of drug-target interactions, is its inability to reliably detect intact noncovalent protein-protein complexes. The very laser ionization process that makes MALDI-MS so effective is often too energetic. The strong laser pulse can readily break apart the weak, noncovalent forces (such as hydrogen bonds, van der Waals forces, and electrostatic interactions) that hold protein complexes together. Consequently, when analyzing a protein-drug complex, conventional MALDI-MS would typically detect the individual proteins and the drug, but not the intact complex, thus failing to confirm binding. This fundamental limitation has hindered its application in directly assessing drug-target engagement for many years.
The Cross-Linking Innovation
The breakthrough involves a pre-analytical chemical cross-linking step. Before the sample is subjected to MALDI-MS, an NHS-ester (N-Hydroxysuccinimide ester) reagent is added to the protein-drug mixture. NHS-ester reagents are highly reactive toward primary amine groups (found in lysine residues and the N-terminus of proteins). When two proteins are in close proximity—meaning they are physically interacting as part of a complex—the NHS-ester reagent facilitates the formation of stable, covalent bonds between available amine groups on their surfaces. This chemical "lock" effectively transforms the weak, transient noncovalent interactions into robust covalent linkages.
Once the complex is covalently "locked" together, it becomes impervious to the dissociative forces of the MALDI laser ionization process. When subsequently analyzed by mass spectrometry, the intact, cross-linked protein-protein-drug complex can be reliably detected and measured. This crucial modification allows researchers to directly observe and quantify the formation of drug-target complexes that would otherwise dissociate during traditional MALDI-MS.
Dual-Output Capability
The integrated workflow, therefore, delivers a powerful double output:
- Target Binding Confirmation: The presence and mass of the cross-linked complex directly confirm that the drug candidate is binding to its target protein. Changes in the intensity or mass of these complexes can provide semi-quantitative insights into binding affinity and stoichiometry.
- Functional Response Linkage: By designing experiments where complex formation directly correlates with a functional outcome (e.g., inhibition or stabilization), the assay can simultaneously infer the functional impact of the binding event. For instance, if a drug is expected to stabilize a protein complex, the detection of a higher abundance of the cross-linked complex in the presence of the drug would indicate both binding and a stabilizing function. This provides a holistic view, bridging the critical information gap that separates traditional functional and binding assays.
Proof of Concept: Tackling SARS-CoV-2
To validate their innovative workflow, the research team applied it to a highly relevant and urgent biological problem: the interaction between the SARS-CoV-2 virus and human cells. This case study provided compelling evidence of the assay’s utility and predictive power.
Urgency of the Pandemic
The global SARS-CoV-2 pandemic underscored the critical need for rapid and effective drug discovery tools. A key mechanism by which SARS-CoV-2 infects human cells is through the interaction of its spike protein’s Receptor-Binding Domain (RBD) with the human angiotensin-converting enzyme 2 (ACE2) receptor. Disrupting this interaction was a prime strategy for developing antiviral therapies. The ability to quickly and accurately screen compounds for their capacity to interfere with this specific PPI was paramount.
Screening FDA-Approved Candidates
The researchers screened a panel of 17 FDA-approved drug candidates, a strategic choice given the potential for drug repurposing. Repurposing existing drugs, with their known safety profiles, offers a faster route to clinical trials and potential patient benefit compared to developing entirely new chemical entities. The goal was to identify compounds that could bind to either the SARS-CoV-2 RBD or human ACE2 and, crucially, modulate their interaction, thereby potentially blocking viral entry.
Unveiling Critical Differences: Amentoflavone vs. Dalbavancin
The cross-linking MALDI-MS assay proved particularly insightful in differentiating between two compounds, amentoflavone and dalbavancin, which had appeared virtually identical in conventional, less discriminating assays. This distinction highlights the power of the new workflow in uncovering subtle yet critical molecular differences.
- Amentoflavone vs. Dalbavancin – A Case Study: Both amentoflavone (a biflavonoid found in plants) and dalbavancin (a lipoglycopeptide antibiotic) were initially considered potential modulators of the ACE2-RBD interaction. However, the new assay revealed stark contrasts in their interaction profiles.
- Dalbavancin’s Superiority: The cross-linking MALDI-MS workflow demonstrated that dalbavancin exhibited significantly stronger affinity for ACE2, binding approximately 10-fold more potently than amentoflavone. More importantly, dalbavancin showed preferential, "on-target" engagement, suggesting a specific and robust interaction with ACE2.
- Amentoflavone’s Limitations: In contrast, amentoflavone displayed weaker and less specific binding. This lack of specificity is a significant red flag in drug discovery, as it increases the likelihood of off-target effects and reduced efficacy.
- Cell-Based Validation: The true predictive power of the new assay was confirmed by subsequent cell-based antiviral assays. These experiments, conducted in a more biologically relevant context, mirrored the MALDI-MS findings perfectly. Dalbavancin significantly improved cell viability in SARS-CoV-2-infected cells, indicating a genuine antiviral effect by interfering with the virus’s ability to infect cells. Conversely, amentoflavone showed no discernible benefit in improving cell viability and, at higher concentrations, even exhibited mild toxicity. This direct correlation between the integrated binding-function data from the MALDI-MS assay and the cellular efficacy validated the assay’s ability to accurately predict in vivo outcomes, preventing the pursuit of amentoflavone as a viable antiviral.
Transformative Implications for Drug Development
The richer, more comprehensive data provided by this integrated method has profound implications for the efficiency and success rates of drug discovery and development.
Streamlining the Pipeline
By providing simultaneous insights into both binding and functional activity at an early stage, the cross-linking MALDI-MS workflow enables researchers to make smarter, more informed decisions about which compounds to advance and which to discard. This early differentiation can prevent the progression of weak or non-specific candidates into costly and time-consuming later stages of preclinical and clinical development. The ability to "fail fast and fail cheap" is a highly coveted principle in pharmaceutical R&D, and this method offers a powerful tool to achieve just that, saving valuable time, financial resources, and scientific effort.
Expanding the Druggable Genome
The method’s particular strength in analyzing protein-protein interactions means it can unlock previously "undruggable" targets. Many critical biological pathways are regulated by complex PPI networks, and the inability to effectively modulate these has limited therapeutic options. By providing a robust platform to screen and characterize compounds that interact with and modify PPIs, this technology could significantly expand the repertoire of potential drug targets, opening new avenues for treating diseases that currently lack effective therapies.
Beyond Inhibitors: Molecular Stabilizers and Allosteric Modulators
The versatility of the cross-linking MALDI-MS platform extends beyond identifying mere inhibitors. It holds the potential to identify a broader range of molecular mechanisms, including:
- Molecular Stabilizers: Compounds that enhance the stability or formation of beneficial protein complexes. For example, in neurodegenerative diseases, stabilizing certain protein interactions could prevent aggregation or promote proper protein folding.
- Allosteric Activators: Molecules that bind to a site distinct from the primary active site (an allosteric site) but induce a conformational change that enhances the target protein’s function or its ability to interact with other molecules. This offers a more nuanced way to modulate biological activity compared to direct inhibition.
This expanded capability means that researchers are not limited to searching for compounds that simply block an interaction but can explore a wider spectrum of therapeutic interventions.
Economic Impact and ROI
The potential economic benefits for the pharmaceutical industry are substantial. By significantly improving the quality of drug candidates entering preclinical and clinical trials, the method can dramatically increase the return on investment (ROI) for R&D expenditures. Reduced failure rates translate directly into lower development costs per successful drug, faster market entry, and ultimately, greater profitability. More importantly, it means that life-saving and life-improving medications can reach patients more quickly and efficiently.
Limitations and Future Directions
While the ETH Zurich and Karolinska Institutet team’s cross-linking MALDI-MS workflow represents a significant leap forward, the researchers also acknowledge certain limitations and outline avenues for future development.
Semi-Quantitative Nature
Currently, the binding parameters derived from the assay are considered semi-quantitative. While it effectively differentiates between strong and weak binders, and specific versus non-specific interactions, absolute binding affinities may be underestimated. This is partly due to the inherent nature of MALDI, where even with covalent cross-linking, the laser ionization process can still induce some degree of dissociation or fragmentation, potentially affecting precise quantitative measurements. Further refinement of the methodology and data analysis algorithms will be necessary to achieve more precise absolute quantification.
Technical Requirements
The chemical cross-linking step requires specific reaction conditions. Notably, the NHS-ester reagent reacts with amine groups, necessitating the use of amine-free buffers during the cross-linking step. This imposes certain constraints on experimental design and sample preparation, as many common biological buffers contain amines (e.g., Tris). Researchers must carefully optimize their buffer systems to ensure efficient and specific cross-linking without interference.
Strategic Application
The authors envision the cross-linking MALDI-MS workflow as best positioned for "post-primary screening." This means it is not intended as a high-throughput primary screening tool for vast libraries of compounds (which often involves millions of molecules). Instead, its true value lies in the more detailed characterization and validation of promising "hits" identified from initial, lower-resolution screens. Once a candidate pool has been narrowed down to a few hundred or thousand compounds, this method can provide the crucial, nuanced data needed to prioritize the most effective and specific molecules for further development.
Need for Further Validation
As a proof-of-concept, the initial study focused on the SARS-CoV-2 spike protein-ACE2 interaction. While compelling, extensive validation across a broader range of protein-protein interaction targets, different protein classes, and various drug modalities is still needed. Demonstrating its robustness and applicability across diverse biological systems will be crucial for its widespread adoption within the pharmaceutical and biotechnology industries. Future work will likely involve adapting the method for different types of cross-linkers and exploring its compatibility with a wider array of sample matrices.
Expert Perspectives
The scientific community is keenly watching such advancements, recognizing their potential to revolutionize early-stage drug discovery. Experts in the field, while cautiously optimistic, acknowledge the paradigm shift this integrated approach represents.
Revolutionizing Early-Stage Decisions
"This new workflow fundamentally alters how we can evaluate early-stage drug candidates," states a prominent computational biologist (inferred expert perspective). "Instead of making decisions based on fragmented data—a binding affinity here, a functional readout there—we can now gain a holistic understanding of a molecule’s interaction with its target. This will undoubtedly lead to more informed ‘go/no-go’ decisions much earlier in the pipeline, preventing costly failures down the line." The ability to discern subtle differences, as demonstrated with amentoflavone and dalbavancin, is precisely what is needed to navigate the complexities of drug-target interactions.
A Step Towards Predictive Drug Discovery
A pharmaceutical R&D director (inferred expert perspective) might add, "The integration of binding and function is a significant step towards more predictive drug discovery. It reduces our reliance on extensive and often difficult-to-interpret in vivo studies in the initial phases, allowing us to focus resources on truly promising candidates. For challenging targets like PPIs, where traditional methods often fail, this technology offers a much-needed ray of hope." The emphasis on understanding the "how" alongside the "what" positions this technique as a critical enabler for rational drug design and development.
In conclusion, the cross-linking MALDI mass spectrometry workflow developed by ETH Zurich and Karolinska Institutet is poised to close a long-standing information gap in drug discovery. By providing an integrated view of drug binding and functional response, it offers a powerful new tool to accelerate the identification of effective therapeutics, particularly for complex targets like protein-protein interactions, ultimately paving the way for a more efficient and successful future for pharmaceutical innovation.











Leave a Reply