In a landmark development for the field of neuroscience, a multidisciplinary team of researchers has unveiled a transformative method for mapping the intricate wiring of the brain. By utilizing molecular "barcodes" made of ribonucleic acid (RNA), scientists can now track thousands of individual neural connections simultaneously, achieving a level of detail and speed that was previously considered unattainable. This breakthrough, led by the University of Illinois Urbana-Champaign, marks a significant shift in connectomics—the study of the brain’s structural and functional connections—and offers a new lens through which to view the progression of neurodegenerative diseases such as Alzheimer’s and Parkinson’s.
The research, published in the prestigious journal Nature Methods, introduces a platform called Connectome-seq. This technology effectively converts the physical architecture of the brain into a massive data-processing problem that can be solved using high-throughput genetic sequencing. By bypassing the labor-intensive requirements of traditional imaging, Connectome-seq allows researchers to reconstruct the "wiring diagram" of the brain with single-synapse resolution, providing a blueprint of how neurons communicate across vast distances.
The Architectural Challenge of the Human Brain
To understand the magnitude of this advancement, one must consider the sheer complexity of the organ in question. The mammalian brain is composed of billions of neurons, each forming thousands of synaptic connections with other cells. These connections, known collectively as the connectome, dictate everything from basic motor functions to complex cognitive processes and emotional regulation.
Historically, mapping these connections has been a Herculean task. For decades, the "gold standard" for brain mapping was electron microscopy (EM). This process involves slicing brain tissue into sections thinner than a human hair, imaging each slice with an electron microscope, and then using advanced software—or sometimes manual labor—to trace the path of every individual axon and dendrite across thousands of images. While EM provides unparalleled spatial resolution, it is notoriously slow and expensive. Mapping a single cubic millimeter of brain tissue can take years of processing time and generate petabytes of data, making it impractical for studying whole-brain circuits or comparing multiple subjects.
"When engineering a computer, you need to know the circuitry of the central processing unit," explained Boxuan Zhao, the study’s lead author and a professor of cell and developmental biology at the University of Illinois Urbana-Champaign. "If you don’t know how everything is wired together, you can’t understand its function, optimize it, or fix it when something breaks. We are approaching the brain the same way."
Connectome-seq: How Molecular Barcoding Works
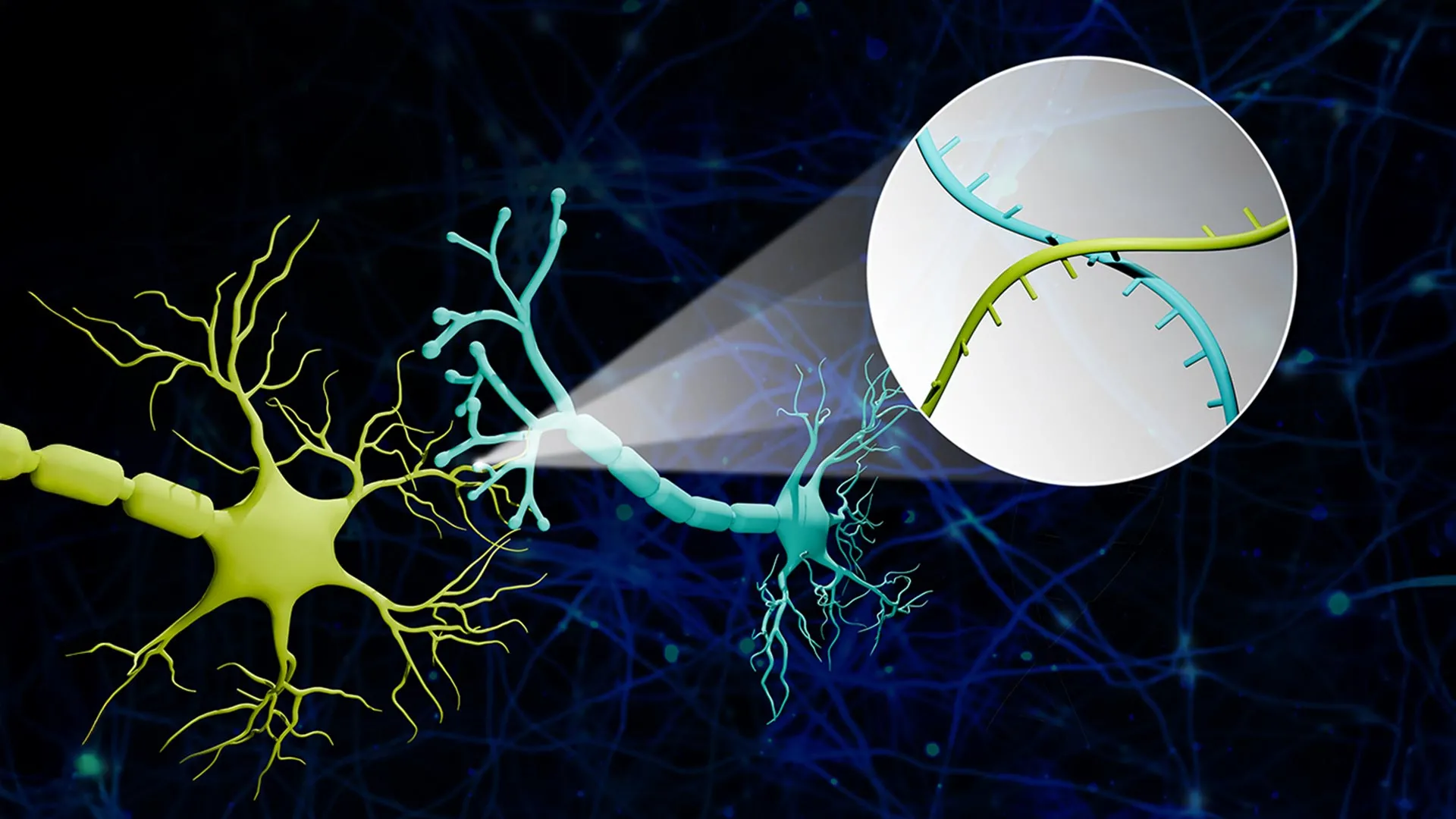
The innovation behind Connectome-seq lies in its ability to label neurons not with light or dyes, but with information. The process begins by assigning a unique RNA "barcode"—a specific sequence of nucleotides—to each individual neuron in a target area. These barcodes act as digital fingerprints for the cells.
To ensure these barcodes reach the points of connection, the researchers engineered specialized proteins designed to transport the RNA sequences from the neuron’s cell body (the soma) down the long, spindly axons to the synapses. The synapse is the microscopic junction where one neuron sends a chemical signal to another. When two neurons are connected, their respective barcodes end up in close proximity at this junction.
The researchers then employ a sophisticated biochemical process to isolate these synapses from the rest of the brain tissue. Once isolated, high-throughput sequencing—the same technology used to sequence the human genome—is used to read the pairs of barcodes found within each synapse.
Zhao utilized a vivid analogy to describe the process: "Imagine a big bunch of balloons. The main body of each balloon has its unique barcode stickers all over it, and some move down to the end of the string. If two balloons are tied together at the end, the two barcodes meet at the junction. Then we snip out the knots and sequence the barcodes in each one. If the same knot has stickers from balloon A and balloon B, we know these two balloons are tied together."
By applying this logic to thousands of neurons simultaneously, the team can reconstruct a sophisticated map representing the connections among vast populations of cells without the need for painstaking visual reconstruction.
Validating the Technology: The Pontocerebellar Circuit
To demonstrate the efficacy of Connectome-seq, the research team focused on a specific mouse brain circuit known as the pontocerebellar circuit. This pathway is essential for motor coordination and sensory integration, linking the pons (part of the brainstem) to the cerebellum.
The team successfully mapped more than 1,000 neurons within this circuit, identifying thousands of specific synaptic connections. The speed of the process was a primary highlight; what would have taken months or years using traditional microscopy was completed in a fraction of the time.
More importantly, the high-resolution data revealed previously unknown patterns of connectivity. The analysis identified direct links between specific cell types that were not previously thought to interact in the adult brain. These findings suggest that current models of brain wiring may be oversimplified and that Connectome-seq could lead to a major revision of neuroanatomical textbooks.
A Timeline of Brain Mapping Evolution
The development of Connectome-seq is the latest milestone in a century-long effort to visualize the brain’s internal workings.
- Late 1800s: Camillo Golgi and Santiago Ramón y Cajal develop the Golgi stain, allowing individual neurons to be seen under a light microscope for the first time.
- 1950s-1960s: The advent of electron microscopy allows scientists to see synapses, the actual gaps between neurons.
- 1986: Scientists complete the first-ever full connectome of a living organism, the roundworm C. elegans, which has only 302 neurons. The project took over a dozen years.
- 2000s: The development of "Brainbow" technology uses fluorescent proteins to color individual neurons in living animals, though it remains limited by the number of distinct colors available.
- 2010s: Viral tracing methods using modified rabies or herpes viruses allow researchers to follow paths from one neuron to its neighbors, but often lack the resolution to identify specific synapses at scale.
- 2024: The introduction of Connectome-seq by Zhao and his colleagues allows for high-throughput, sequencing-based mapping with single-synapse precision.
Implications for Alzheimer’s and Neurodegenerative Research
The most significant impact of Connectome-seq may lie in the realm of clinical medicine. Neurodegenerative diseases like Alzheimer’s are characterized by the progressive loss of synapses—a process that often begins years or even decades before the death of the neurons themselves and the onset of cognitive symptoms.
Current diagnostic tools, such as MRI or PET scans, can show brain shrinkage or the buildup of toxic proteins (like amyloid-beta or tau), but they cannot visualize the microscopic breakdown of individual circuits. Connectome-seq provides a platform to study these "weak links" in the neural chain.
"With sequencing-based approaches, the time and cost are greatly reduced, which really makes it possible to see differences in different brains," Zhao noted. "We could see where connections change, where the most vulnerable parts of the brain are, perhaps before symptoms even appear."
By comparing the connectomes of healthy brains with those at various stages of Alzheimer’s, researchers can pinpoint exactly which circuits fail first. This "circuit-guided" approach could lead to highly targeted therapies. For instance, if a specific set of connections in the hippocampus is identified as the "kick-start" for the disease’s progression, clinicians could theoretically use deep brain stimulation or localized drug delivery to strengthen those specific synapses, potentially slowing or halting the disease.
Expert Reactions and Future Directions
The scientific community has reacted with cautious optimism to the UIUC study. While the technology currently focuses on mouse models, the scalability of sequencing suggests it could eventually be applied to human brain tissue samples obtained post-mortem.
Dr. Elena Martinez, a neuroscientist not involved in the study, commented on the potential for data integration. "The beauty of Connectome-seq is that it can be combined with other ‘omics’ data. We can see not only which neurons are connected but also what genes those specific neurons are expressing at the time. This gives us a functional and structural profile simultaneously."
However, challenges remain. While sequencing tells us which cells are connected, it does not inherently provide the physical location of those connections in the 3D space of the brain. Zhao’s lab is already working on integrating spatial transcriptomics with Connectome-seq to bridge this gap, creating a map that is both genetically informative and spatially accurate.
The ultimate goal, according to Zhao, is to map the entire mouse brain—a feat that would involve billions of connections. "With improvements already underway in our lab, we are confident that we can make it even better and eventually reach the goal of mapping the whole mouse brain," he said.
Economic and Strategic Impact
The shift from microscopy-based mapping to sequencing-based mapping also has significant economic implications for the research sector. DNA sequencing costs have plummeted over the last two decades, following a trajectory even steeper than Moore’s Law in computing. By piggybacking on this existing infrastructure, connectomics can move from a "Big Science" endeavor restricted to a few elite institutions with multi-million dollar microscopes to a more accessible field for laboratories worldwide.
The research was supported by a Neuro-omics Initiative grant from the Wu Tsai Neurosciences Institute at Stanford University, as well as funding from the Elsa U. Pardee Foundation and the Edward Mallinckrodt Jr. Foundation. These investments reflect a growing consensus that the next frontier of medicine lies in understanding the "circuitry of the soul."
As Connectome-seq continues to evolve, it promises to provide the detailed "wiring diagram" necessary to not only understand the healthy human mind but to repair it when it falters. By translating the mysteries of the brain into the language of genetic sequences, Zhao and his team have opened a new chapter in the quest to understand the most complex structure in the known universe.











Leave a Reply