In an era defined by data proliferation, the ability to process and analyze information with speed and precision is paramount for data scientists and analysts. While the Python pandas library has long been the cornerstone for data manipulation, a common observation is that many practitioners, especially those new to the field, often adopt suboptimal coding practices. These habits, frequently learned through introductory tutorials, can lead to technically functional but inefficient, hard-to-read, and difficult-to-maintain codebases. Industry experts now underscore the critical importance of transitioning from basic pandas usage to more advanced patterns to unlock significant performance gains, enhance code clarity, and foster robust data workflows. This shift is not merely about aesthetic preference but represents a fundamental evolution in how data operations are executed, directly impacting project scalability, resource utilization, and team collaboration.
The journey of a data scientist with pandas typically begins with intuitive methods, such as iterating through rows using iterrows(), employing numerous intermediate variables, and executing repetitive merge() calls. While these approaches offer immediate results and facilitate initial learning, they often conceal underlying inefficiencies that become pronounced when dealing with large datasets—a common scenario in today’s data-intensive environments. As organizations increasingly rely on data-driven insights, the demand for high-performance computing in routine data tasks has grown exponentially. This necessitates a move towards vectorized operations and optimized data structures that leverage pandas‘ C-level underpinnings, rather than relying on slower Python-level iterations. The evolution of best practices in pandas reflects a broader maturation within the data science community, moving from ad-hoc scripting to engineering-grade data pipelines.
Streamlining Operations with Method Chaining
One of the most impactful transformations in pandas coding style is the adoption of method chaining. This pattern allows a sequence of data transformations to be written as a single, contiguous expression, thereby significantly improving readability and reducing cognitive load. Instead of declaring multiple intermediate DataFrame objects, which can clutter a script and make it harder to follow the data’s journey, method chaining creates a clear, sequential flow. This approach minimizes the creation of unnecessary variables, each representing a snapshot of the data at a particular stage, often adding "noise" to the codebase without providing unique identifiers that warrant their existence.
Consider a common scenario: filtering, dropping null values, deriving a new column, and then sorting the data. A novice approach might involve four distinct lines of code, each assigning the result to a new variable (df1, df2, df3, result). While functional, this verbosity obscures the overall data flow. Method chaining condenses this into a single, fluid expression, typically wrapped in parentheses for multi-line readability. A crucial element in chaining is the use of lambda functions within methods like assign(). This allows the current state of the DataFrame within the chain to be referenced, preventing NameError or stale references that can arise from attempting to access a DataFrame by a variable name defined outside the current scope of the chain. Furthermore, the practice of using inplace=True within chained operations is strongly discouraged. Such methods return None, immediately breaking the chain and leading to unpredictable behavior or errors. Avoiding in-place operations also enhances code predictability and debugging, as each step in the chain explicitly returns a modified DataFrame.
Encapsulating Complexity with the pipe() Pattern

While method chaining excels at sequencing built-in pandas operations, data transformation pipelines often require custom logic that is too complex or specific to be inline. This is where the pipe() method becomes indispensable. The pipe() pattern allows data scientists to integrate custom functions into a method chain, preserving the chain’s flow and readability. It acts as a bridge, passing the DataFrame as the first argument to any callable function, enabling the encapsulation of complex transformations within named, testable functions.
For instance, a data cleaning pipeline might involve a custom normalization function that scales specific columns. Without pipe(), this would necessitate breaking the chain, applying the function, and then restarting the chain, disrupting the elegant flow. With pipe(), the normalization function, complete with its internal logic, can be seamlessly inserted into the chain. This not only keeps the code organized but also promotes modularity. Each piped function can be developed and tested independently, a significant advantage for debugging and maintaining large, intricate data processing pipelines. Moreover, pipe() contributes to self-documenting code. By dividing a complex pipeline into distinct, labeled functions, the purpose of each step becomes immediately clear from the function name, without requiring readers to delve into the implementation details. This enhances collaborative development and makes it easier to modify or even temporarily skip steps during development or debugging, simply by commenting out a pipe() call without affecting the rest of the chain.
Optimizing Data Integration: Efficient Joins and Merges
Data integration, often performed using pandas.DataFrame.merge(), is a cornerstone of data analysis. However, it is also one of the most frequently misused functions, leading to significant performance bottlenecks and data integrity issues. The two most common pitfalls are silent row inflation due to many-to-many joins and the failure to identify unjoined rows. When merge() encounters duplicate values in the join key on both sides of the merge, it performs a Cartesian product, which can explode a DataFrame from hundreds of rows to millions without an explicit error. This "silent row inflation" is a critical data quality issue, producing seemingly correct but vastly oversized results.
The validate parameter in merge() is a powerful safeguard against such issues. By specifying validate='many_to_one', one_to_one', or one_to_many', practitioners can enforce assumptions about the uniqueness of their join keys. If these assumptions are violated, pandas immediately raises a MergeError, preventing downstream analytical errors. This proactive validation is crucial for maintaining data integrity in complex pipelines. Equally important for debugging is the indicator=True parameter. This adds a special _merge column to the result, indicating whether each row originated from the "left_only," "right_only," or "both" DataFrame. This immediate feedback mechanism is invaluable for quickly identifying rows that failed to match, particularly in left or right joins where unmatched rows might otherwise silently disappear or appear as nulls. For cases where dataframes share an index, pandas.DataFrame.join() offers a performance advantage over merge(), as it operates directly on the index without the overhead of searching through specified columns.
Advanced Groupby Operations with transform()
The groupby() method is fundamental for aggregating data, but its full potential is often underutilized. While agg() is commonly used to return a single row per group, transform() offers a distinct advantage for scenarios requiring group-level statistics to be added back to the original DataFrame without altering its shape. Unlike agg(), which collapses groups, transform() returns a Series or DataFrame with the same index and shape as the original, where each row is filled with its corresponding group’s aggregated value.

This capability is particularly powerful for tasks such as standardizing data within groups or appending group averages as new columns. For example, calculating the average revenue per customer segment and adding it as a new column to every row in the original DataFrame can be achieved in a single, efficient step using transform(). A manual approach would involve an agg() operation followed by a separate merge() call, incurring additional computational cost and complexity. transform() bypasses the need for explicit merging, as pandas handles the alignment internally, leading to significant performance gains, especially on large datasets. Another critical optimization for groupby() is the observed=True argument when dealing with categorical columns. Without observed=True, pandas can compute results for all possible categories defined in the column’s dtype, even those not present in the actual data. This can lead to unnecessary computations and empty groups, particularly problematic with high-cardinality categorical columns. Specifying observed=True ensures that computations are performed only on categories that are actually present in the data, enhancing efficiency.
Vectorized Conditional Logic for Performance
One of the most common performance pitfalls in pandas is the overuse of apply() with lambda functions for row-wise conditional logic. While intuitive, this approach is inherently slow because it iterates over DataFrame rows at the Python level, bypassing pandas‘ optimized, C-level vectorized operations. For binary conditions, NumPy‘s np.where() is the direct and vastly superior replacement. It allows for conditional assignment based on a boolean array, executing at C-speed and offering dramatic performance improvements.
For more complex scenarios involving multiple conditions, np.select() provides a clean and highly efficient solution. This function maps directly to an if/elif/else structure, evaluating a list of conditions in order and assigning the corresponding choice from a parallel list. The default parameter ensures that rows not meeting any specified condition receive a fallback value. Benchmarks consistently show np.select() to be 50 to 100 times faster than an equivalent apply() operation on large datasets. Beyond explicit conditional logic, pandas offers specialized functions for numeric binning: pd.cut() for equal-width bins and pd.qcut() for quantile-based bins. These functions automatically handle bin labeling and edge cases, returning a categorical column without any need for manual conditional assignments, further demonstrating pandas‘ commitment to vectorized efficiency.
Identifying and Avoiding Performance Pitfalls
Beyond the explicit patterns for improvement, data scientists must also be vigilant about common pandas anti-patterns that significantly degrade performance. The iterrows() method, while seemingly straightforward for row-by-row processing, is a prime example. It iterates over DataFrame rows as (index, Series) pairs, building a complete Series object for each row and executing Python code on it sequentially. This overhead makes iterrows() notoriously slow—up to 100 times slower than vectorized alternatives for a DataFrame with 100,000 rows. The Pandas community strongly advises against iterrows() and itertuples() for anything but the smallest datasets or when no vectorized alternative exists. In almost all cases, np.where(), np.select(), or groupby() operations can replace such loops with far greater efficiency.
Similarly, apply(axis=1), though often slightly faster than iterrows(), still suffers from the same fundamental problem: executing Python-level code for each row. Whenever a task can be expressed using built-in NumPy or pandas functions, these vectorized methods will invariably outperform apply(axis=1). Another often-overlooked source of sluggishness lies in object dtype columns. When pandas stores strings or mixed types as object dtype, operations on these columns revert to slower Python-level processing. For columns with low cardinality (e.g., status codes, region names), converting them to a categorical dtype can yield significant speedups for groupby() and value_counts() operations, as categorical data is stored more efficiently and processed at C-level.
Finally, "chained assignment" is a pervasive anti-pattern that can lead to ambiguous behavior and SettingWithCopyWarning messages. An expression like df[df['revenue'] > 0]['label'] = 'positive' attempts to modify a potentially temporary DataFrame copy, leading to undefined behavior where the original DataFrame may or may not be updated. The unambiguous and correct way to perform conditional assignment is by using .loc with a boolean mask: df.loc[df['revenue'] > 0, 'label'] = 'positive'. This ensures that the assignment is performed directly on the original DataFrame, avoiding unexpected outcomes and warnings.
Conclusion and Broader Implications
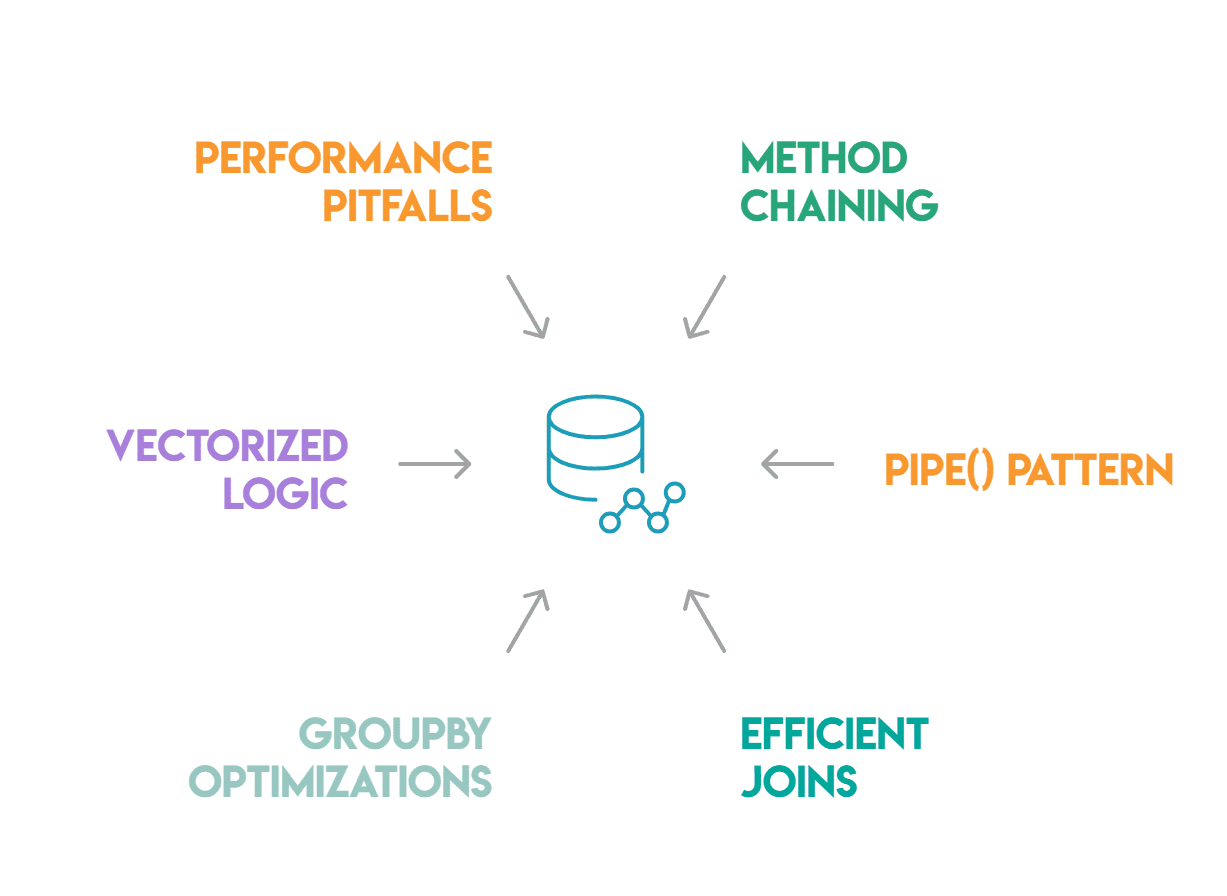
The mastery of these advanced pandas patterns—method chaining, the pipe() pattern, efficient joins, groupby() optimizations, vectorized conditional logic, and the avoidance of common performance pitfalls—marks a critical distinction between code that merely functions and code that performs optimally. These practices are not academic exercises but pragmatic necessities for data scientists working with real-world datasets that demand efficiency, readability, and maintainability. By embracing these methodologies, practitioners can develop robust data pipelines that are not only faster and consume fewer computational resources but are also easier to understand, debug, and extend by team members.
The implications extend beyond individual projects. Organizations that foster a culture of adopting these advanced pandas techniques stand to benefit from reduced computational costs, faster time-to-insight, and higher data quality. The cumulative effect of numerous small inefficiencies—a slow loop here, an unvalidated merge there, an unoptimized object dtype column—can quietly undermine entire data initiatives, leading to missed deadlines, increased infrastructure spending, and diminished trust in data-driven decisions. As Nate Rosidi, a data scientist, adjunct professor, and founder of StrataScratch, emphasizes, these issues accumulate quietly because they rarely cause outright failures, making them persistent challenges. Addressing them systematically, one pattern at a time, represents a tangible pathway to elevating the overall maturity and effectiveness of data science workflows across the industry. The ongoing evolution of pandas itself, coupled with the growing complexity of data challenges, necessitates that data professionals continually refine their skills to remain at the forefront of efficient and responsible data stewardship.












Leave a Reply