The pharmaceutical industry faces an escalating crisis of escalating costs, protracted timelines, and persistently high failure rates in bringing new medicines to market. With typical timelines for successful drugs stretching from 10 to 15 years, and often longer, and the cost of development ranging from hundreds of millions to multiple billions of dollars per drug, the economic pressures are immense. Compounding this challenge, the inflation-adjusted cost of drug development has approximately doubled every nine years, a phenomenon dubbed "Eroom’s Law" – the inverse of Moore’s Law, highlighting a worrying trend of diminishing returns despite advancements in technology. At the core of this inefficiency lies a fundamental problem: a significant proportion of promising drug candidates fail in clinical trials due to a lack of efficacy, indicating a profound disconnect between preclinical assessments and actual biological outcomes in humans.
The Unyielding Challenge of Drug Discovery: A Deep Dive into Eroom’s Law
Drug discovery and development is an inherently complex and capital-intensive endeavor. It begins with target identification and validation, followed by lead discovery and optimization, preclinical testing, and finally, a multi-phase clinical trial process before regulatory approval. Each stage presents unique scientific and logistical hurdles. The initial stages involve screening vast libraries of compounds against specific biological targets or cellular phenotypes, a process that has historically been labor-intensive and slow. Despite significant technological leaps, including high-throughput screening and genomics, the efficiency of drug development has paradoxically declined. Eroom’s Law illustrates this stark reality: for every billion dollars spent on R&D, the number of new drugs approved by regulatory bodies like the FDA has halved roughly every nine years since 1950. This inverse correlation underscores a systemic challenge where innovation struggles to keep pace with the increasing complexity of biological systems and regulatory requirements.
The financial implications are staggering. A 2016 study by the Tufts Center for the Study of Drug Development estimated the cost to develop a new prescription drug that gains market approval at $2.6 billion, a figure that continues to rise. This cost includes both out-of-pocket expenses and the time cost of capital. A major driver of these exorbitant costs is the high attrition rate, particularly in clinical trials. Each failure represents not just a lost opportunity but also the squandering of significant resources, driving up the overall cost for the few drugs that eventually succeed.
The Efficacy Gap: Why Preclinical Assays Fall Short
A critical reason for the high clinical failure rate, particularly in Phase II and Phase III trials, is the fundamental lack of efficacy. An extensive analysis of clinical trial data between 2010 and 2017 revealed that 40% to 50% of drugs that fail in clinical trials do so because they simply do not work as intended in human subjects. This stark statistic underscores a profound limitation in conventional preclinical assays: their inability to accurately predict a drug’s biological activity and therapeutic potential in a complex living system.
The core of this problem lies in what can be termed the "snapshot assay problem." Traditional drug discovery methods often rely on assays that reduce complex, dynamic cellular behavior into static, single-point measurements. These assays typically capture a cell’s state at a single moment in time, often after a prolonged exposure to a compound, failing to account for the intricate, time-evolving responses of biological systems. For instance, common assays for cytotoxicity or receptor binding provide a binary or quantitative readout at a specific endpoint, missing the nuanced progression of cellular changes that define a drug’s mechanism of action or its off-target effects.
Furthermore, many conventional techniques, such as transcriptome profiling, involve lysing (destroying) cells to extract their molecular components. While invaluable for understanding gene expression at a population level, this destructive nature makes it impossible to track the dynamic gene expression patterns of an individual cell over multiple time points. This limitation means researchers lose critical information about how a cell adapts, recovers, or succumbs to a drug’s influence, insights that are crucial for predicting long-term efficacy and potential side effects. The result is an incomplete, fragmented picture of drug-cell interactions, poorly reflecting the true biological complexity.
AI’s Promise and Present Limitations in Drug Discovery
In recent years, the field of artificial intelligence (AI) in drug discovery has attracted immense investment and considerable optimism, positioned as a potential panacea for many of the industry’s woes. Between 2024 and 2025 alone, the sector saw an impressive 612 venture rounds, accumulating approximately $19.9 billion in total capital, according to a sector review by DealForma. Proponents argue that AI can accelerate target identification, optimize compound design, predict toxicity, and even identify new indications for existing drugs by sifting through vast datasets far more efficiently than human researchers.
However, despite this substantial investment and technological promise, the clinical validation of AI’s impact remains limited and mixed. To date, AI has not demonstrably improved the stubbornly high 90% clinical failure rate for drug candidates. While AI has certainly enhanced efficiency in certain preclinical stages, such as virtual screening and lead optimization, its ability to fundamentally alter the success rate in human trials is yet to be proven. The challenge lies in the quality and relevance of the data fed into AI models, and the inherent difficulty in translating in silico or in vitro predictions into in vivo human outcomes. If the underlying experimental data suffers from the "snapshot assay problem" and other limitations, even sophisticated AI algorithms may struggle to make accurate predictions about efficacy in humans.
Phenotypic Screening: A Renewed Hope with Hurdles
The limitations of target-centric drug discovery, where researchers focus on modulating a single, well-defined molecular target, have led to a resurgence of interest in phenotypic drug discovery. This approach shifts the focus from a specific molecular target to observing complex cellular or organismal behavior in response to a compound. Instead of asking "which enzyme does this drug inhibit?", phenotypic screening asks "what does this drug do to the cell?" This allows for the discovery of drugs with novel mechanisms of action, or those that act on multiple targets simultaneously (polypharmacology), which can be particularly beneficial for complex diseases.
Historically, many blockbuster drugs, including penicillin and aspirin, were discovered through phenotypic screening long before their molecular targets were fully understood. The renewed interest stems from the recognition that many diseases are multifactorial, and modulating a single target may not be sufficient for therapeutic success. By observing the overall cellular response, researchers hope to identify compounds that elicit desirable physiological changes, regardless of their precise molecular interactions.
However, phenotypic approaches come with their own set of challenges. These include "hit validation" – confirming that an observed phenotype is robust and reproducible; "target deconvolution" – the often-difficult task of identifying the specific molecular targets responsible for the observed phenotype; and the general difficulty in translating broad phenotypic signals into precise mechanistic insights. Without understanding the underlying mechanism, optimizing a drug candidate or predicting its potential side effects can be significantly harder, making the path to clinical development more arduous.
Live Cell Dynamics: A Paradigm Shift in Preclinical Assessment
It is within this landscape of persistent challenges that Live Cell Dynamics (LCD) emerges as a potentially transformative technology. Developed by scientists at Soley Therapeutics, LCD is a self-supervised machine learning pipeline designed to extract crucial dose- and time-dependent cellular state information directly from continuous brightfield images, crucially without the need for traditional stains or labels. This innovative method was detailed in a January 2026 Scientific Reports paper, marking a significant step towards addressing the limitations of static, destructive assays.
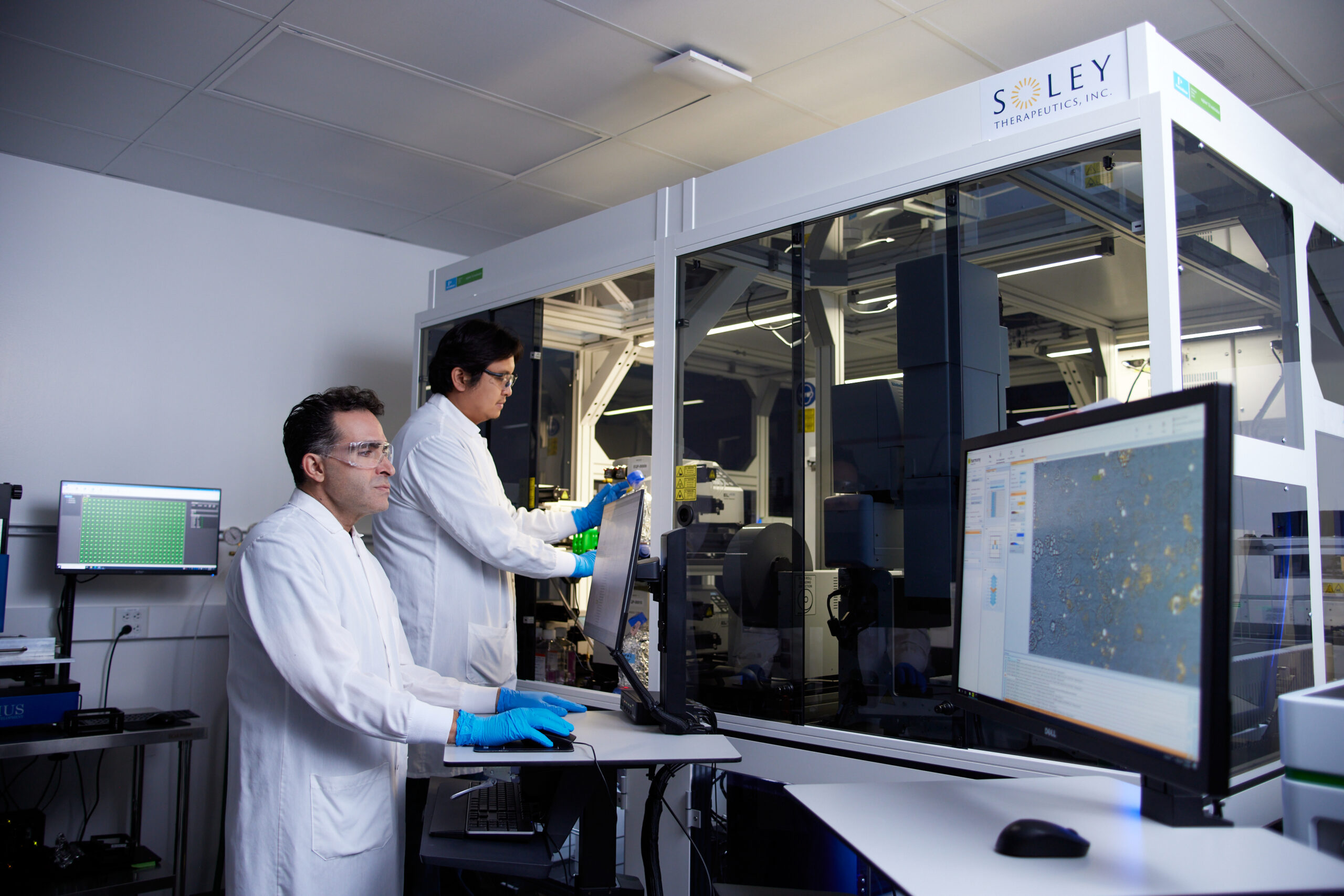

Kurosh Ameri, co-founder and CSO of Soley Therapeutics, explained the fundamental shift LCD represents: "By treating cellular response as time-resolved information rather than a static snapshot, LCD enables mechanism classification, compound comparison, and detection of complex biology through measurable trajectories." He further elaborated that this approach provides "early forward-looking biological signal rather than a late binary readout, shifting drug discovery from observing damage to forecasting a drug’s direction and future impact." This ability to observe and analyze the process of cellular response, rather than just the endpoint, is a critical departure from conventional methods.
Unpacking the Soley Therapeutics Study: Methodology and Key Findings
The study conducted by Soley Therapeutics employed a robust methodology to demonstrate LCD’s capabilities. Researchers pre-trained the machine learning model on 189 compounds and subsequently evaluated its performance on 81 additional "held-out" compounds, ensuring an unbiased assessment. These compounds spanned 10 distinct mechanisms of action and were tested using a single, well-characterized human osteosarcoma cell line (U2OS), a common model in cancer research.
The results were compelling. LCD significantly outperformed traditional methods, specifically cell count and CellProfiler-based feature extraction, in detecting phenotypic activity across all tested doses and time points. The most pronounced advantages were observed at early time points and lower doses, where conventional methods often struggle to discern subtle biological changes. This early detection capability is vital, as it allows researchers to identify promising compounds much earlier in the discovery pipeline, potentially saving time and resources.
Crucially, the study found that incorporating multiple doses and time points incrementally improved the mechanism-of-action classification. This multi-dimensional data allowed LCD to disentangle mechanisms that might appear similar or indistinguishable at later, more advanced stages of cellular response, when damage or overwhelming effects might mask the primary mode of action. Ameri highlighted this, stating, "Learned representations from LCD preserved signal in those early regimes and performed strongly across dose and time, while the CellProfiler baseline tended to be comparable only later, or lower at early time points." This suggests LCD can provide a more nuanced and accurate understanding of how drugs interact with cells from the very outset.
Detecting Polypharmacology and Overcoming Technical Hurdles
One of the most significant advantages demonstrated by LCD is its ability to detect polypharmacology – the phenomenon where many drugs affect multiple biological targets simultaneously. While common and often therapeutically beneficial, polypharmacology is notoriously difficult to identify using conventional methods, typically requiring extensive and costly assay panels. The Soley Therapeutics study showed that using only brightfield imaging, the LCD model successfully flagged both Aurora kinase and JAK inhibitor activity. This finding was consistent with prior studies that had required extensive kinome profiling, a much more resource-intensive technique, to reach the same conclusion. This capability could dramatically streamline the characterization of complex drug candidates, providing a more complete picture of their biological footprint.
However, working with brightfield imaging presents its own unique set of challenges. As Ameri noted, "Brightfield is difficult because the signal is subtle, not evident to the naked eye, contrast is low, and small changes in optics, focus, plate position, or day-to-day setup can create batch effects that swamp biology." To overcome these inherent difficulties, the paper outlines two ingenious training innovations:
- Plane-agnostic augmentation: This technique teaches the model to recognize underlying biological changes irrespective of minor variations in focal plane or imaging artifacts. By training the model to be robust to these optical inconsistencies, it can focus on true biological signals.
- Cross-batch sampling: This innovation forces the model to learn features that are stable and consistent across different experimental runs and batches. By minimizing the influence of technical noise and day-to-day variations, the model can effectively separate genuine biological signals from experimental artifacts.
These methodological advancements are critical to making LCD a reliable and scalable tool in drug discovery. The results collectively demonstrate that "LCD can represent compound behavior as a profile across dose and time, not a single label. Those profiles contain enough structure to separate closely related mechanisms and expose mixed activity, which is exactly the kind of complexity that shows up in development," Ameri concluded.
Implications for Drug Development: A Forward-Looking Signal
The implications of Live Cell Dynamics for the drug discovery pipeline are profound. By providing an early, dynamic, and comprehensive biological signal, LCD has the potential to:
- Improve Efficacy Prediction: Moving beyond static snapshots, LCD offers a richer, time-resolved understanding of drug action, potentially leading to more accurate predictions of clinical efficacy and significantly reducing the failure rate in human trials.
- Accelerate Lead Optimization: The ability to rapidly characterize complex mechanisms of action and detect polypharmacology with simple brightfield imaging can dramatically accelerate the lead optimization phase, guiding medicinal chemists towards compounds with more desirable profiles.
- Reduce Costs: By identifying ineffective or problematic compounds earlier, LCD could prevent the significant investment of resources into candidates destined for clinical failure, thereby contributing to a reduction in the overall cost of drug development.
- Enhance Mechanism of Action Elucidation: The detailed temporal profiles generated by LCD can provide deeper insights into how drugs interact with cells, even for phenotypically discovered compounds where the target might initially be unknown. This can bridge the gap between phenotypic screening and target deconvolution.
- Facilitate Personalized Medicine: As LCD is expanded to patient-derived cells, it could potentially offer a platform for predicting individual patient responses to drugs, paving the way for more effective personalized treatment strategies.
The Road Ahead: Validation and Broader Application
Despite the promising results, the study acknowledges important limitations and outlines clear next steps. The current research utilized a single, well-characterized cancer cell line (U2OS) under controlled laboratory conditions. This means LCD’s performance in more complex and biologically relevant models, such as primary cells, patient-derived organoids (3D tissue models), or animal disease models, remains to be thoroughly investigated. The central question left open by the work is whether the performance advantages observed in a controlled compound library will hold up across the "messier, more heterogeneous biology of disease models."
According to Soley Therapeutics, the immediate next step is to expand LCD to additional cell types, including primary cells and those directly relevant to human diseases. This will involve broadening the coverage of mechanisms of action and integrating LCD into prospective use within active drug discovery programs. For LCD to truly deliver on its promise and gain widespread adoption in the pharmaceutical industry, it will need to be rigorously validated in settings that more closely mimic human disease conditions. Only after such validation can definitive claims about its clinical impact be fairly evaluated.
Broader Industry Impact and Expert Perspectives
The introduction of technologies like Live Cell Dynamics represents a critical juncture for the pharmaceutical industry. Industry observers suggest that such innovations are essential to overcome the current productivity crisis. If successfully validated across a wider range of disease models, LCD could become an indispensable tool, complementing existing high-throughput screening methods and AI platforms. It could foster collaborations between technology developers and pharmaceutical companies, driving a more data-rich and mechanistically informed approach to drug discovery.
The potential for LCD to identify polypharmacology early, for example, is particularly intriguing. Many successful drugs exert their effects through multiple pathways, and the ability to detect and understand these complex interactions without extensive, expensive follow-up assays could significantly accelerate the development of multi-modal therapies for complex diseases like cancer, neurodegenerative disorders, and autoimmune conditions.
Ultimately, the journey from a promising preclinical technology to a clinically impactful tool is long and fraught with challenges. However, by directly addressing the long-standing problem of preclinical-to-clinical disconnect and offering a dynamic, label-free window into cellular behavior, Live Cell Dynamics offers a compelling vision for a more efficient, cost-effective, and ultimately more successful future for drug discovery. The scientific community and pharmaceutical industry will be closely watching Soley Therapeutics’ subsequent validation efforts as they seek to move this innovative technology closer to impacting human health.








Leave a Reply