The pharmaceutical industry grapples with an intractable problem: the escalating cost and protracted timelines of drug development, coupled with persistently high rates of clinical failure. Bringing a new drug from initial discovery to market approval typically consumes between 10 to 15 years, and often even longer, demanding an investment that can range from hundreds of millions to multiple billions of dollars. This financial burden is not static; alarming trends indicate that the inflation-adjusted cost of drug development has approximately doubled every nine years, a phenomenon often referred to as "Eroom’s Law"—Moore’s Law in reverse, reflecting a decline in research and development efficiency over time. This dire economic landscape underscores an urgent need for transformative innovations in the drug discovery pipeline.
A primary driver behind these staggering statistics is the pervasive issue of drug efficacy, or rather, the lack thereof, in clinical trials. A comprehensive analysis of clinical trial data spanning from 2010 to 2017 revealed a sobering truth: a significant proportion, between 40% and 50%, of drugs that fail in clinical trials do so because they simply do not work as intended. This critical shortfall highlights a fundamental disconnect between preclinical assays and the complex biological realities they are meant to model, indicating that current early-stage testing methods poorly predict a drug’s ultimate performance in living systems.
Against this backdrop, the field of AI-driven drug discovery has emerged as a beacon of hope, attracting substantial investment and widespread optimism. In 2024 and 2025 alone, the sector witnessed 612 venture rounds, accumulating an impressive $19.9 billion in total capital, according to a sector review by DealForma. Yet, despite this influx of resources and advanced computational power, AI has, to date, not demonstrably altered the grim 90% clinical failure rate that plagues drug development. While AI promises to accelerate target identification, compound synthesis, and lead optimization, its impact on the fundamental problem of predicting clinical efficacy remains a challenge still largely unmet.
The "Snapshot Assay" Problem: A Core Limitation
One of the most significant contributors to the pervasive clinical trial failure rate lies in the inherent structural limitations of conventional drug discovery assays. For decades, standard laboratory methods have simplified complex, dynamic cellular behaviors into static, single-point measurements. These "snapshot assays" capture a cell’s state at one specific moment, often after a predetermined incubation period, failing to account for the intricate, time-dependent cascades of biological events that unfold in response to a therapeutic agent. This reductionist approach fundamentally misrepresents the evolving nature of cellular biology, where a drug’s effect is not a single outcome but a trajectory of interactions over time.
Furthermore, many traditional assays, particularly those involving high-throughput screening or detailed molecular profiling like transcriptome analysis, necessitate the destruction of cells. While invaluable for identifying specific molecular signatures, this destructive nature makes it impossible to track the dynamic gene expression or phenotypic changes of an individual cell or cell population across multiple time points. Researchers are left with fragmented data, unable to observe how a cell’s response develops, adapts, or changes course over hours or days, thereby obscuring crucial mechanistic insights that could differentiate between effective and ineffective compounds.
These limitations have spurred a renewed interest in phenotypic drug discovery (PDD), an approach that shifts the focus from modulating a single, pre-identified molecular target to observing complex cellular behavior in response to a compound. PDD aims to identify molecules that induce a desired biological effect, such as halting cancer cell proliferation or restoring neuronal function, without necessarily knowing the precise molecular target upfront. This strategy inherently embraces the complexity of biology, moving beyond the "one gene, one drug, one disease" paradigm that has often proven too simplistic for multifaceted conditions.
However, phenotypic drug discovery, while promising, is not without its own set of significant challenges. Historically, these approaches have struggled with "hit validation"—confirming that an observed phenotypic effect is robust and reproducible. More critically, they face difficulties in "target deconvolution," the often arduous process of identifying the specific molecular targets or pathways through which a compound exerts its beneficial effects. Without this mechanistic understanding, translating phenotypic signals into actionable insights for drug optimization and understanding potential off-target effects remains a formidable hurdle, hindering the progression of promising candidates.
Live Cell Dynamics: A Paradigm Shift with Machine Learning
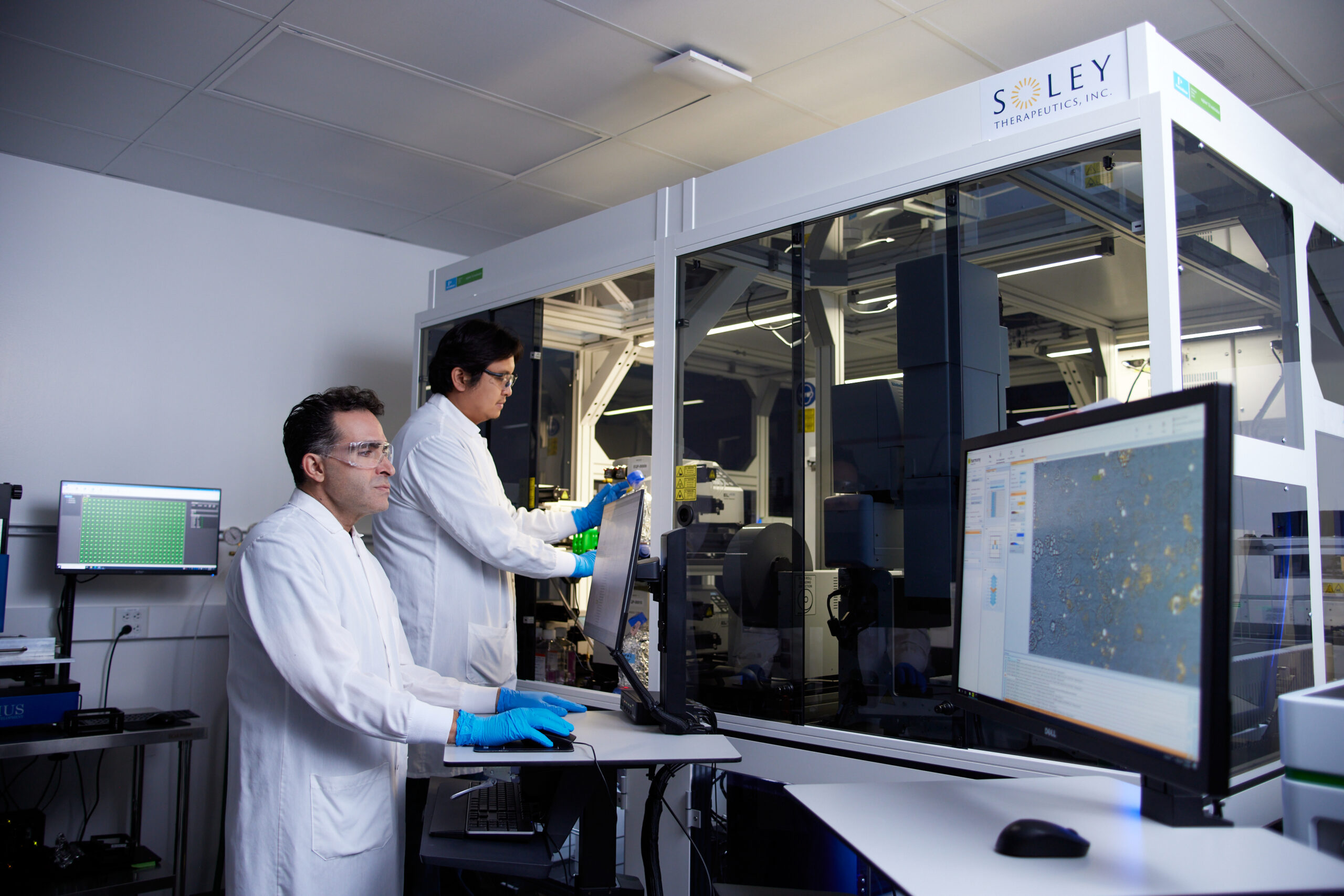
In response to these deeply entrenched challenges, a groundbreaking methodology known as Live Cell Dynamics (LCD) has emerged, promising to redefine how drugs are discovered and validated. LCD represents a self-supervised machine learning pipeline specifically designed to extract rich, dose- and time-dependent cellular state information directly from continuous brightfield images, crucially without the need for cumbersome or perturbing stains or labels. This innovative method was detailed in a seminal paper published in January 2026 in Scientific Reports by a team of scientists from Soley Therapeutics.
Kurosh Ameri, co-founder and Chief Scientific Officer of Soley Therapeutics, elucidated the core philosophy behind LCD: "By treating cellular response as time-resolved information rather than a static snapshot, LCD enables mechanism classification, compound comparison, and detection of complex biology through measurable trajectories." He further emphasized the profound implications of this shift, stating, "This provides early forward-looking biological signal rather than a late binary readout, shifting drug discovery from observing damage to forecasting a drug’s direction and future impact." This approach moves beyond merely detecting cell death or a static endpoint, allowing researchers to observe the process of cellular response, predicting whether a drug’s trajectory is leading towards desired therapeutic outcomes or undesirable side effects.
The study conducted by Soley Therapeutics demonstrated the power of LCD through rigorous evaluation. Researchers pre-trained the machine learning model on a dataset of 189 compounds and subsequently assessed its performance on 81 additional, held-out compounds, encompassing 10 distinct mechanisms of action. All experiments were conducted using a single, well-characterized human osteosarcoma cell line (U2OS), providing a controlled environment for initial validation.
The results were compelling: LCD significantly outperformed traditional methods, including simple cell count and CellProfiler-based feature extraction, in detecting phenotypic activity across all tested doses and time points. The advantages of LCD were particularly pronounced at early time points and lower doses, regimes where conventional assays often fail to capture subtle but crucial biological signals. This early detection capability is vital, as it allows for quicker identification of promising candidates and earlier deselection of ineffective ones, saving considerable time and resources downstream. Furthermore, the study highlighted that incorporating multiple doses and time points incrementally improved the accuracy of mechanism-of-action classification, enabling the model to disentangle complex biological mechanisms that might appear similar in their late-stage effects but differ fundamentally in their initial response pathways.

Ameri elaborated on these findings, noting, "Learned representations from LCD preserved signal in those early regimes and performed strongly across dose and time, while the CellProfiler baseline tended to be comparable only later, or lower at early time points." This superior performance in early detection is critical for drug discovery, as it provides a much-needed window into the initial cellular responses, which are often indicative of a drug’s true mechanism and potential efficacy.
Unmasking Polypharmacology with Label-Free Imaging
One of the most challenging aspects of drug discovery is the phenomenon of polypharmacology, where a single drug can affect multiple biological targets simultaneously. While often seen as a complication, judicious polypharmacology can sometimes be therapeutically beneficial, particularly in complex diseases like cancer or neurodegenerative disorders. However, identifying and characterizing polypharmacology conventionally requires extensive, costly, and often label-dependent assay panels, making it notoriously difficult to detect and understand early in the discovery process.
Remarkably, the LCD model, using only simple brightfield imaging—a label-free, non-invasive technique—successfully flagged both Aurora kinase and JAK inhibitor activity. This detection was consistent with prior studies that had necessitated extensive kinome profiling, a much more resource-intensive and expensive method, to reach the same conclusions. This capability of LCD to uncover complex, multi-target interactions from basic visual data represents a significant leap forward, offering a more efficient and cost-effective way to identify and characterize polypharmacological effects early in drug development.
The subtle nature of brightfield imaging signals, however, presents its own set of technical hurdles. "Brightfield is difficult because the signal is subtle, not evident to the naked eye, contrast is low," Ameri explained. "And small changes in optics, focus, plate position, or day-to-day setup can create batch effects that swamp biology." These technical variations can easily obscure the genuine biological signals, making robust and reproducible data extraction a significant challenge.
To overcome these inherent difficulties, the Soley Therapeutics paper outlined two key training innovations embedded within the LCD pipeline. The first is "plane-agnostic augmentation," a technique that teaches the machine learning model to recognize genuine biological phenomena rather than focusing on artifacts introduced by variations in focus plane. The second innovation, "cross-batch sampling," forces the model to learn features that are stable and consistent across different experimental runs, effectively separating true biological signal from technical noise and batch-to-batch variability. These advancements are crucial for ensuring the reliability and generalizability of the LCD platform.
Ameri concluded by summarizing the broader implications of these findings: "The results demonstrate that LCD can represent compound behavior as a profile across dose and time, not a single label. Those profiles contain enough structure to separate closely related mechanisms and expose mixed activity, which is exactly the kind of complexity that shows up in development." This ability to generate nuanced, multi-dimensional profiles of compound activity moves beyond simplistic "yes/no" readouts, providing a richer, more predictive understanding of how drugs interact with biological systems.
Implications and Future Directions
The advent of Live Cell Dynamics holds profound implications for the future of drug discovery. By providing early, dynamic, and mechanism-rich insights, LCD has the potential to significantly accelerate the identification of promising drug candidates, reduce attrition rates in preclinical stages, and ultimately lower the overall cost and timeline of drug development. Its ability to detect complex polypharmacology and differentiate subtle mechanistic variations could lead to the development of more targeted and effective therapies, especially for diseases where single-target approaches have fallen short. The non-invasive nature of brightfield imaging also offers practical advantages, making it amenable to high-throughput screening and continuous monitoring without compromising cell viability.
However, the study also acknowledged important limitations and outlined critical next steps. The initial validation of LCD was conducted using a single, well-characterized cancer cell line (U2OS) under highly controlled laboratory conditions. While providing a robust proof-of-concept, this controlled environment does not fully replicate the complexity and heterogeneity of human physiology. Consequently, whether the performance advantages observed in this specific context will translate to more biologically relevant models—such as primary cells, patient-derived organoids, or complex disease models—remains an open and central question. The "messier, more heterogeneous biology of disease models" presents a formidable challenge that will require further rigorous testing.
According to Soley Therapeutics, the immediate future plans involve expanding the LCD platform to encompass additional cell types, including various primary cell cultures and disease-relevant models that more closely mimic human pathology. Furthermore, efforts will be directed towards broadening the platform’s mechanism coverage to analyze an even wider array of biological processes and drug targets. Crucially, the technology will need to undergo prospective use in active drug programs, moving beyond retrospective analysis to demonstrate its predictive power in real-world drug development scenarios. Only through such comprehensive validation in settings closer to human disease can claims about LCD’s transformative clinical impact be fairly and fully evaluated.
In essence, Live Cell Dynamics represents a significant advancement in the application of machine learning to fundamental biological problems. By moving beyond static observations to embrace the dynamic nature of cellular life, it offers a promising pathway to address the long-standing efficacy problem in drug discovery. As the technology matures and undergoes further validation in diverse and complex biological systems, it has the potential to reshape how pharmaceutical research is conducted, bringing safer, more effective, and more efficiently developed medicines to patients worldwide.








Leave a Reply