In a landmark advancement for the field of neuroscience, researchers have unveiled a transformative methodology for charting the intricate architecture of the brain, utilizing molecular "barcodes" to map neural connectivity with unprecedented speed and precision. Developed by a multidisciplinary team led by the University of Illinois Urbana-Champaign, this new platform, dubbed Connectome-seq, allows scientists to bypass the labor-intensive and time-consuming bottlenecks of traditional imaging. By converting the physical wiring of the brain into a high-throughput sequencing problem, the researchers have successfully mapped thousands of neural connections within the mouse brain, revealing previously unknown pathways and setting the stage for a new era in the study of neurodegenerative diseases such as Alzheimer’s.
The human brain, containing approximately 86 billion neurons and trillions of synaptic connections, remains one of the most complex structures in the known universe. Understanding how these cells communicate is fundamental to deciphering the mechanics of thought, memory, and behavior. However, the sheer scale of the "connectome"—the comprehensive map of neural connections—has historically outpaced the capabilities of available technology. The introduction of Connectome-seq represents a paradigm shift, offering a scalable solution that can simultaneously resolve thousands of individual synapses, a feat that has remained elusive for decades.
The Evolution of Connectomics: From Slicing to Sequencing
To appreciate the significance of Connectome-seq, it is necessary to examine the historical context of brain mapping. For over a century, the primary method for studying neural circuits involved histology and microscopy. In the late 19th century, Camillo Golgi and Santiago Ramón y Cajal used silver staining to visualize individual neurons, a breakthrough that earned them the Nobel Prize. However, these methods only provided a glimpse into the brain’s structure.
In the modern era, the "gold standard" for connectomics has been serial section electron microscopy (EM). This process involves slicing brain tissue into sections thinner than a human hair, imaging each slice with an electron microscope, and then using advanced computer algorithms or manual labor to reconstruct the three-dimensional paths of axons and dendrites. While EM offers unmatched resolution, it is notoriously slow. Mapping a single cubic millimeter of brain tissue can take years of processing and petabytes of data storage.
While more recent innovations, such as light-sheet microscopy and viral tracing, have accelerated the process, they often lack the resolution to confirm actual synaptic connections. Viral tracers might show that a neuron "projects" to a certain region, but they cannot always identify which specific cell it is talking to. Connectome-seq addresses this "resolution gap" by utilizing the language of genetics—RNA sequencing—to identify direct synaptic partners.
The Mechanics of Connectome-seq: Molecular Barcoding Explained
The innovation behind Connectome-seq lies in its ability to assign a unique identity to every neuron. The research team, led by Boxuan Zhao, a professor of cell and developmental biology at the University of Illinois Urbana-Champaign, engineered a system where each neuron produces a unique RNA "barcode." This barcode acts as a molecular fingerprint, distinguishing one cell from thousands of its neighbors.
The process functions through a series of sophisticated biological steps:
- Labeling: Each neuron is tagged with a unique RNA sequence.
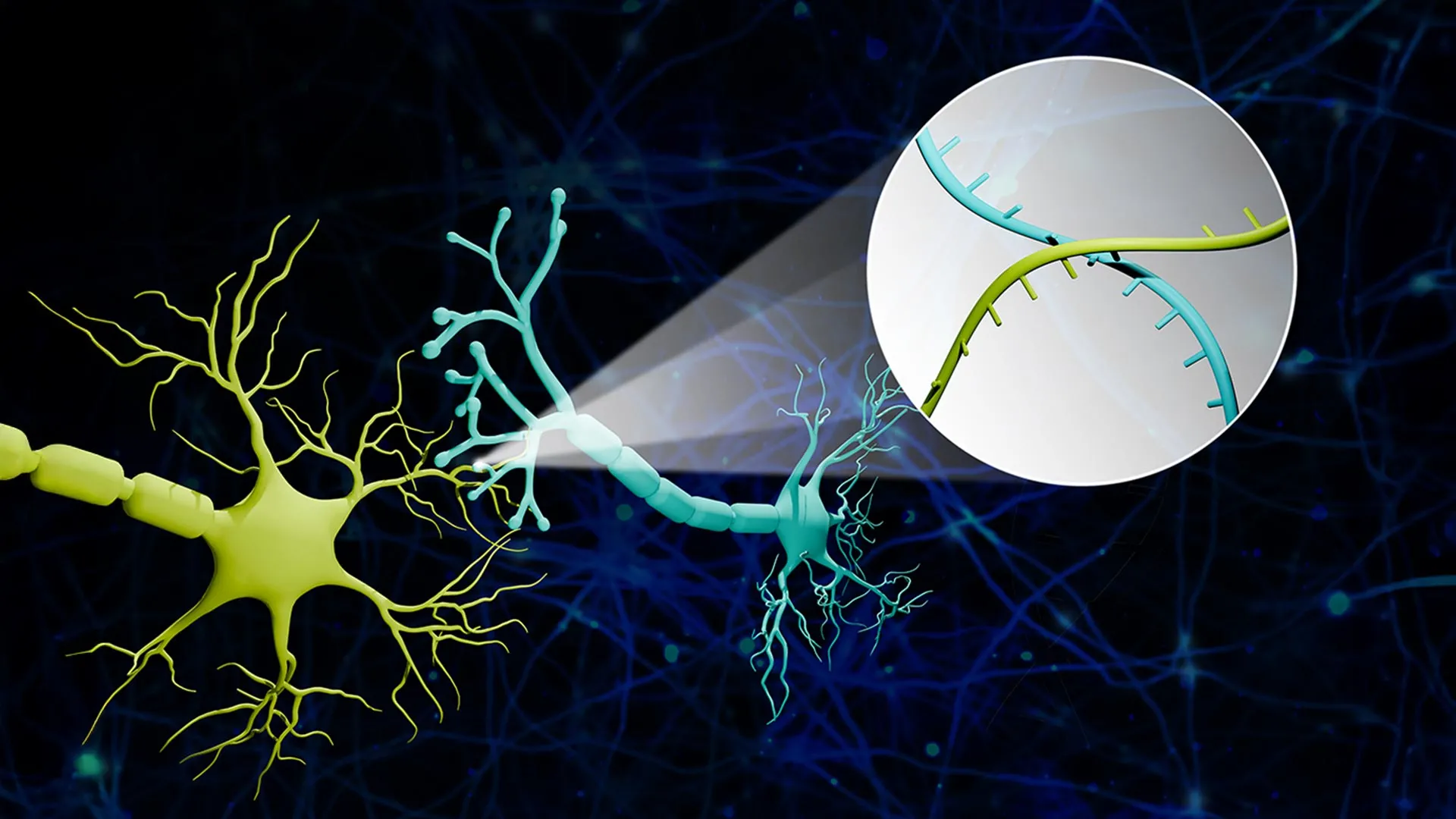
- Transport: Specialized proteins, acting as molecular shuttles, carry these RNA barcodes from the cell body (soma) down the long, spindly axon to the synapse—the junction where one neuron meets another.
- Synaptic Capture: At the synapse, the barcodes from the "sender" neuron and the "receiver" neuron are brought into close proximity.
- Sequencing: Researchers isolate these synaptic junctions and use high-throughput RNA sequencing to read the pairs of barcodes found together.
Professor Zhao utilized a vivid analogy to describe this complex process: "Imagine a big bunch of balloons. The main body of each balloon has its unique barcode stickers all over it, and some move down to the end of the string. If two balloons are tied together at the end, the two barcodes meet at the junction. Then we snip out the knots and sequence the barcodes in each one. If the same knot has stickers from balloon A and balloon B, we know these two balloons are tied together."
By applying this logic to the brain, the team can reconstruct a sophisticated map of connectivity based on the frequency and pairing of these molecular "knots."
Validating the Model: New Discoveries in the Pontocerebellar Circuit
To demonstrate the efficacy of Connectome-seq, the researchers targeted the pontocerebellar circuit in the mouse brain. This circuit is a critical pathway linking the pons (part of the brainstem) to the cerebellum, and it plays a vital role in motor coordination, sensory integration, and cognitive functions.
The team successfully mapped more than 1,000 neurons within this circuit. The results, published in the journal Nature Methods, were surprising even to the researchers. The high-resolution data revealed connectivity patterns that had never been documented in the adult mouse brain. Specifically, the analysis identified direct links between certain cell types that were previously thought to be independent or only indirectly connected.
These findings suggest that our current understanding of brain circuitry—even in well-studied animal models—is incomplete. The ability of Connectome-seq to reveal these "hidden" connections in a matter of weeks, rather than years, underscores its potential to rewrite neuroscience textbooks.
Comparative Advantages: Speed, Cost, and Scalability
The primary advantage of sequencing-based connectomics over imaging-based methods is scalability. In traditional imaging, the time required to map a circuit increases exponentially with the volume of the tissue. In contrast, sequencing costs and timelines are decreasing rapidly due to advancements in the genomics industry.
According to data extrapolated from the study, Connectome-seq offers several key improvements:
- Temporal Efficiency: What once took months of manual reconstruction can now be processed in a fraction of the time using automated sequencing machines.
- Single-Synapse Resolution: Unlike traditional bulk labeling, which might show a general "highway" of neurons, Connectome-seq identifies the specific "house addresses" (synapses) where connections occur.
- Simultaneity: The platform can track thousands of connections at once, providing a holistic view of a network rather than a piecemeal look at individual cells.
Professor Zhao emphasized that this approach treats the brain like a computer’s central processing unit (CPU). "If you don’t know how everything is wired together, you can’t understand its function, optimize it or fix it when something breaks," he noted. By turning brain wiring into a digital sequencing problem, the researchers have created a tool that can be used by laboratories worldwide without the need for multi-million-dollar electron microscopy suites.
Implications for Alzheimer’s and Neurodegenerative Disease Research
Perhaps the most significant application of Connectome-seq lies in the study of neurodegenerative and psychiatric disorders. Diseases like Alzheimer’s, Parkinson’s, and schizophrenia are increasingly viewed as "connectopathies"—conditions where the primary pathology is the disruption of neural circuits.
In Alzheimer’s disease, for example, the loss of synapses often occurs long before the death of neurons and the onset of cognitive decline. Current diagnostic tools, such as MRI or PET scans, lack the resolution to see these early synaptic changes. Connectome-seq could allow researchers to compare the "wiring diagrams" of healthy brains against those in the early stages of disease.
"We could see where connections change, where the most vulnerable parts of the brain are, perhaps before symptoms even appear," Zhao explained. This capability opens the door to "circuit-guided therapeutic interventions." If scientists can identify the specific "weak link" that initiates the cascade of degeneration, they may be able to develop treatments that specifically strengthen those connections, potentially slowing or halting the progression of the disease.
Collaborative Support and Future Horizons
The development of Connectome-seq was a collaborative effort, supported by prestigious institutions and foundations. The research received a Neuro-omics Initiative grant from the Wu Tsai Neurosciences Institute at Stanford University, reflecting the interdisciplinary nature of the project which blends biology, engineering, and data science. Additional funding was provided by the Elsa U. Pardee Foundation and the Edward Mallinckrodt Jr. Foundation.
Looking ahead, the research team is already working on enhancing the platform. Current goals include increasing the number of unique barcodes available to map even larger populations of neurons and improving the efficiency of the molecular "shuttles" that carry the barcodes to the synapses.
"With improvements already underway in our lab, we are confident that we can make it even better and eventually reach the goal of mapping the whole mouse brain," Zhao said.
The ultimate ambition of the field is to apply these techniques to human brain tissue, perhaps through the use of brain organoids or post-mortem samples. While mapping the human connectome remains a monumental task, the success of Connectome-seq provides a credible roadmap for achieving what was once considered impossible. By translating the physical complexity of the brain into the digital language of DNA sequencing, researchers have provided the scientific community with a powerful new lens through which to view the mysteries of the mind.










Leave a Reply