Evolution, the relentless engine of nature, has long served as the ultimate biological engineer. Within the intricate molecular factories of cells, a constant flux of DNA, RNA, and protein variations arises. Natural selection, the discerning hand of this process, invariably favors those organisms whose molecular machinery operates with the greatest efficiency and adaptability. Humans, in their own quest for optimization, have recognized and harnessed this fundamental principle for millennia. The dawn of agriculture marked humanity’s first foray into influencing evolution, as early farmers meticulously selected the most robust crops and prolific livestock, ensuring that advantageous traits were passed down through generations. This foundational understanding of inheritance and selection paved the way for modern scientific endeavors.
Today, this ancient principle is amplified and refined in the laboratory through a sophisticated methodology known as directed evolution. Scientists, mirroring nature’s selective pressures, engineer novel proteins with tailored functions. These engineered proteins, particularly enzymes and antibodies, are instrumental across a vast spectrum of applications, from life-saving medicines and the large-scale manufacturing of industrial goods to the ubiquitous presence of enzymes in everyday products such as laundry detergents. The ability to precisely alter protein function has revolutionized numerous scientific and commercial fields.
The Limitations of Conventional Directed Evolution
Despite its profound success, traditional directed evolution, while powerful, is not without its inherent constraints. A primary limitation lies in its typical reliance on a constant selection pressure. This approach overwhelmingly favors proteins that maintain a high level of activity continuously. However, this paradigm diverges significantly from the nuanced reality of biological systems. Many proteins do not operate in a static state; rather, they function as dynamic regulators. They act as molecular signals, intricate switches, or "logic gates"—proteins that process multiple incoming signals to make binary, yes-or-no decisions. Crucially, these proteins must be able to transition between states in response to fluctuating environmental conditions.
Consider a protein that briefly activates, then deactivates, only to re-engage later. When evolutionary experiments are designed to reward only a single, sustained state of activity, other essential functional states can inadvertently degrade. This can lead to proteins that lose their capacity for precise switching, a deficiency that can be detrimental, even lethal, to a cell. The challenge of engineering proteins with complex, multi-state behaviors has, therefore, remained a significant hurdle for conventional directed evolution techniques. The inability to reliably replicate the dynamic, context-dependent nature of protein function in living organisms has limited the scope and sophistication of engineered biological systems.
A Novel Light-Based Strategy for Protein Evolution: Optovolution Emerges
Addressing this critical gap, researchers spearheaded by Sahand Jamal Rahi at EPFL’s Laboratory of the Physics of Biological Systems have unveiled a groundbreaking new approach termed "optovolution." This innovative method leverages the precise control offered by light to guide the evolution of proteins capable of performing dynamic functions. Furthermore, optovolution facilitates the development of proteins that can execute simple computational tasks, adhering to the fundamental rules of binary logic.
The seminal study, published in the esteemed journal Cell, represents a significant stride towards aligning laboratory-directed evolution with the intricate operational principles of natural cellular systems. In living organisms, the timing of molecular events and the seamless switching between different functional states are as vital to cellular health and function as the inherent strength or potency of a signal itself. This research underscores the importance of temporal dynamics in protein engineering.
Engineering Yeast Cells: A Biological Crucible for Optimal Protein Selection
To construct their revolutionary system, the research team employed the well-established model organism Saccharomyces cerevisiae, commonly known as budding yeast. This organism holds a dual significance, being indispensable in both the brewing industry and cutting-edge scientific research. The scientists ingeniously redesigned the yeast cell cycle, fundamentally linking cell division to the dynamic behavior of the protein undergoing evolution. For the yeast cell to survive and propagate, the protein in question was required to exhibit a clean and timely transition between its active and inactive states.
The core of their innovation involved connecting the protein’s output signal to a critical regulator that governs the cell cycle. This regulator plays an indispensable role during a specific phase of the cell cycle but becomes toxic if it remains active during another. Consequently, if the engineered protein remained perpetually "on" or "off" for an extended period, the yeast cell would either halt its progression through the cell cycle or perish. Only those yeast cells harboring proteins that successfully switched states at the precise moment dictated by the engineered system were able to continue dividing and proliferate. This created a powerful selective pressure that inherently favored proteins with superior dynamic switching capabilities.
Harnessing Light for Real-Time Evolutionary Control
Light emerged as the key to achieving exquisite control over this complex evolutionary process. The researchers adeptly employed optogenetics, a powerful technique that enables the activation or deactivation of specific genes through the application of light. By delivering precisely timed pulses of light, the scientists could effectively compel the protein under investigation to alternate between its active and inactive states.
Each cycle of the yeast cell’s life, lasting approximately 90 minutes, served as a rapid, high-throughput test of the protein’s switching performance. Proteins that demonstrated optimal dynamic behavior allowed the yeast cell to survive and reproduce, while variants exhibiting suboptimal switching patterns were systematically eliminated from the population. This automated selection process, driven by light and cell viability, enabled optovolution to identify and propagate proteins with enhanced dynamic characteristics without the need for laborious manual screening or iterative experimental adjustments. The efficiency and precision of this light-guided evolution significantly accelerated the discovery of proteins with desired dynamic properties.
Generating Novel Protein Variants and Expanding Color Sensitivity
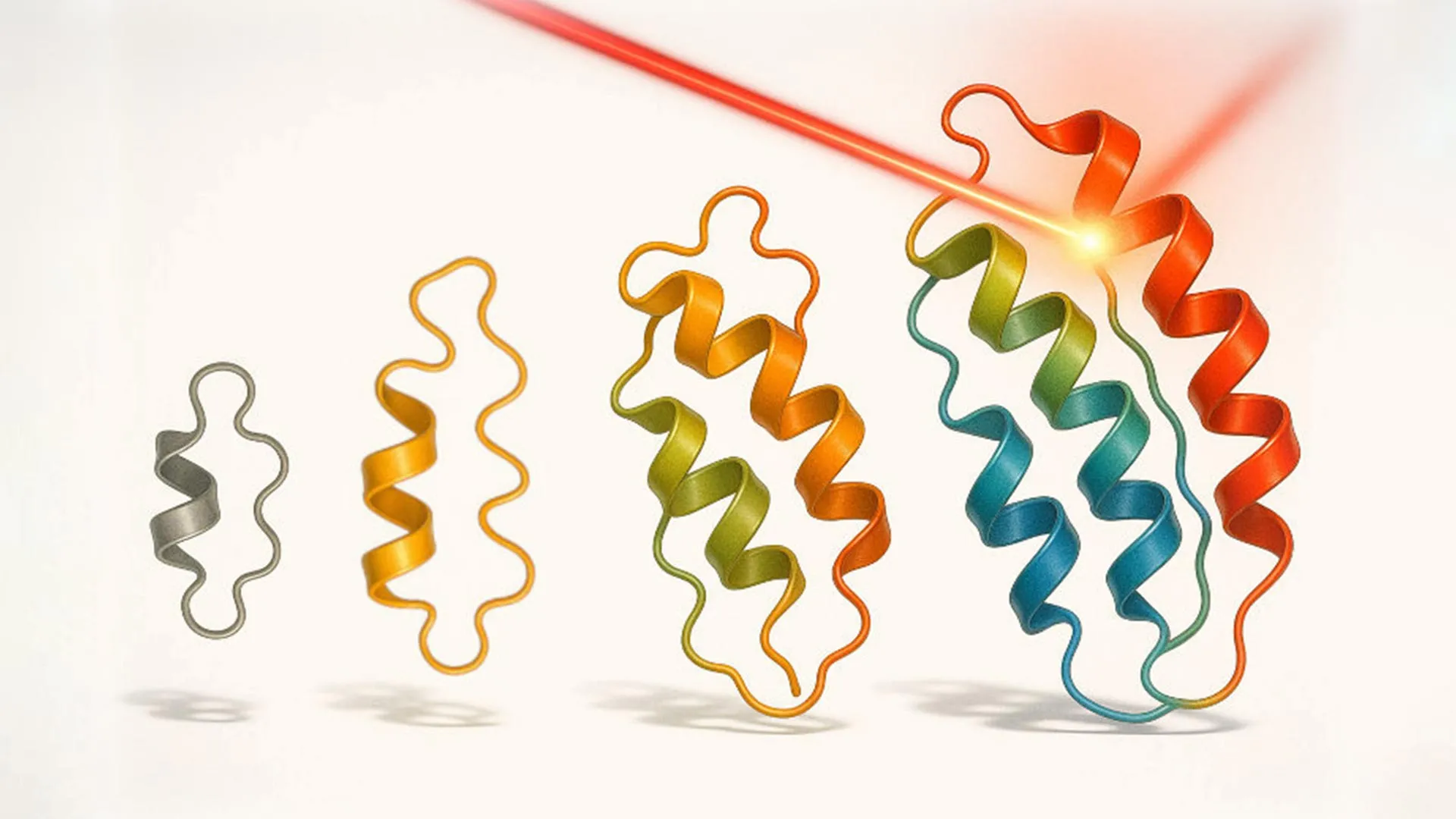
Utilizing the optovolution platform, the research team successfully engineered several distinct classes of proteins. Their initial efforts focused on enhancing a commonly used light-controlled transcription factor. Through repeated cycles of evolution, they generated an impressive 19 novel variants of this transcription factor. These engineered proteins exhibited remarkable improvements, including heightened sensitivity to light stimuli, significantly reduced activity in the absence of light (darkness), and the novel ability to respond to green light, a departure from the blue light sensitivity of the original protein. The development of proteins responsive to "warmer" colors of light, such as green and red, has historically been a formidable challenge in protein engineering. This difficulty stems from the fundamental way these proteins absorb light energy, making them less amenable to manipulation by longer wavelengths. The success in engineering green light sensitivity represents a significant breakthrough in this area.
In a subsequent advancement, the scientists engineered a red light-activated optogenetic system. A key benefit of this new system was its ability to function within yeast cells without the requirement of an externally added chemical cofactor. The evolutionary process fortuitously produced a mutation that inactivated a native yeast transport protein. This unexpected yet beneficial alteration enabled the optogenetic system to utilize light-sensitive molecules that were already intrinsically present within the yeast cell. This discovery significantly simplified the experimental setup and enhanced the practical utility of the optogenetic tools.
Engineering Proteins That Function as Miniature Biological Computers
The versatility of optovolution extended beyond light-sensing proteins, as demonstrated by its application in engineering a transcription factor that operates akin to a single-protein computer. This engineered protein was designed to activate gene expression only when two distinct input signals were simultaneously present: one originating from a light stimulus and the other from a specific chemical signal. This capability to integrate multiple inputs and perform logical operations at the protein level opens up new avenues for creating sophisticated cellular circuits.
The ability of proteins to exhibit dynamic behavior is fundamental to a wide array of biological processes. These include the sensing of environmental changes, the intricate decision-making pathways within cells, and the precise control of cell division. By enabling these critical dynamic behaviors to evolve organically within living cells, optovolution offers transformative possibilities for the fields of synthetic biology, biotechnology, and fundamental biological research.
The implications of this research are far-reaching. Scientists may now be better equipped to design more intelligent and responsive cellular circuits, enabling the development of novel therapeutic agents and biosensors. The creation of optogenetic tools that can independently respond to different colors of light promises greater precision and multiplexing capabilities in experimental manipulations. Furthermore, optovolution provides a powerful new lens through which to investigate the fundamental mechanisms by which complex protein behaviors emerge through the crucible of natural selection. Understanding these evolutionary pathways can lead to a deeper comprehension of cellular function and dysfunction, potentially paving the way for new strategies to combat diseases. The ability to engineer proteins that integrate multiple signals and exhibit dynamic responses could lead to the development of "smart" drugs that activate only under specific cellular conditions, minimizing off-target effects and enhancing therapeutic efficacy. The potential for creating cellular systems that can perform complex computations could revolutionize areas like diagnostics, where cells could be engineered to detect specific disease markers and trigger a response.
The research team is actively exploring the expansion of optovolution to other model organisms and the engineering of more complex computational logic gates within single proteins. This ongoing work promises to further blur the lines between engineered and natural biological systems, ushering in an era of unprecedented control and design capabilities in the life sciences. The journey from understanding nature’s evolutionary blueprint to actively directing it in the laboratory has taken a significant leap forward with the advent of optovolution.












Leave a Reply