Researchers at the University of Illinois Urbana-Champaign have unveiled a transformative approach to mapping the intricate architecture of the brain, utilizing molecular "barcodes" to track how individual neurons connect at the synaptic level. This new technology, dubbed Connectome-seq, represents a significant leap forward in the field of connectomics—the comprehensive study of the brain’s structural and functional connections. By translating the physical wiring of the brain into a high-throughput sequencing problem, the team has successfully charted thousands of neural connections in the mouse brain with a level of speed and precision that was previously unattainable.
The study, led by Boxuan Zhao, a professor of cell and developmental biology, and published in the journal Nature Methods, addresses one of the most persistent challenges in modern neuroscience: the sheer complexity and scale of neural networks. The human brain contains approximately 86 billion neurons, each forming thousands of synapses, while the mouse brain, a primary model for medical research, contains roughly 70 million neurons. Mapping these connections has historically been a labor-intensive process, often taking years or even decades to complete for even small sections of brain tissue. Connectome-seq aims to bypass these bottlenecks, offering a scalable platform that could eventually lead to a complete map of the mammalian brain.
The Evolution of Brain Mapping: From Silver Stains to Sequencing
To appreciate the magnitude of the Connectome-seq breakthrough, it is necessary to examine the history of brain mapping. For over a century, neuroscientists have struggled to visualize the "wiring diagram" of the brain. In the late 19th century, Camillo Golgi and Santiago Ramón y Cajal used silver staining techniques to visualize individual neurons for the first time. While revolutionary, these methods were limited to looking at a tiny fraction of cells in a given sample.
In more recent decades, Electron Microscopy (EM) became the gold standard for mapping synapses. EM provides incredible resolution, allowing scientists to see the physical junctions between neurons. However, the process is agonizingly slow. To map a significant portion of a brain, researchers must slice tissue into sections thinner than a human hair, take millions of high-resolution images, and then use complex algorithms or manual labor to trace the path of each neuron across those slices. The first complete connectome ever mapped was that of the roundworm C. elegans, which has only 302 neurons. That project took more than a dozen years to complete.
While newer light microscopy and sequencing-based tools have emerged, they often suffer from a "resolution gap." Many current tools can identify where a neuron sends its long-reaching fibers (axons), but they cannot definitively confirm which specific cell those fibers connect to. Connectome-seq bridges this gap by focusing on the synapse itself—the precise point of chemical and electrical communication between two cells.
Decoding Connectome-seq: How Molecular Barcoding Works
The innovation behind Connectome-seq lies in its ability to convert physical biological structures into digital data. The process begins by assigning a unique RNA "barcode" to each neuron. These barcodes are essentially short, synthetic sequences of genetic material that act as a unique identification tag for every individual cell.
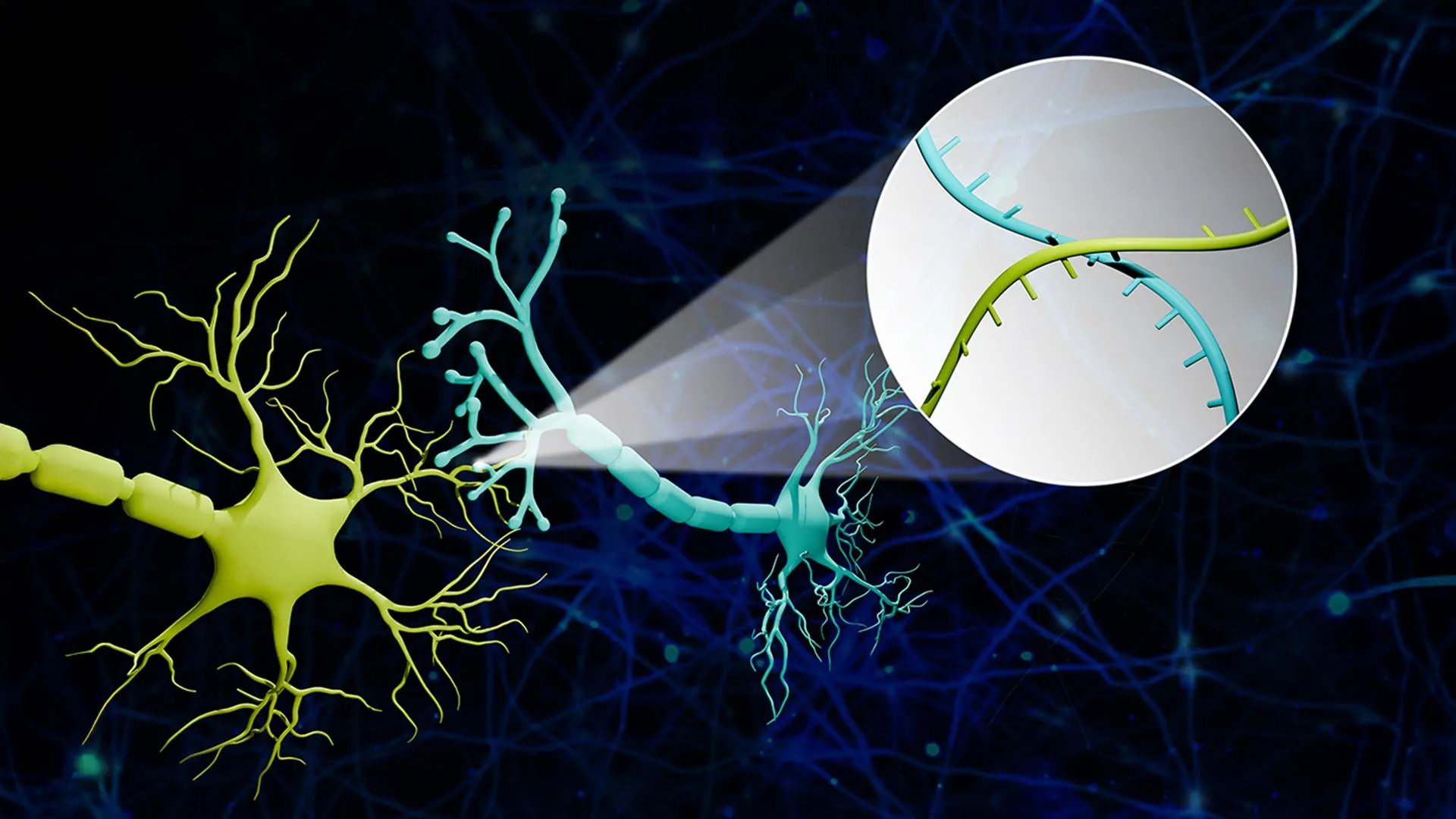
"We translated the neural connectivity problem into a sequencing problem," explained Professor Zhao. To illustrate the concept, Zhao uses an analogy of a massive cluster of balloons. "Imagine a big bunch of balloons. The main body of each balloon has its unique barcode stickers all over it, and some move down to the end of the string. If two balloons are tied together at the end, the two barcodes meet at the junction."
In the brain, the "balloons" are the cell bodies of neurons, and the "strings" are the axons. Specialized proteins are engineered to carry the RNA barcodes from the cell body down the axon to the synapse. When two neurons form a connection, their respective barcodes are brought into close proximity at the synaptic junction. Researchers then isolate these synapses and use high-throughput sequencing—the same technology used for rapid DNA testing and genomic research—to read which barcode pairs are found together.
If a sequence reveals barcode A and barcode B in the same "knot" or synapse, the researchers know with certainty that Neuron A and Neuron B are connected. This method allows for the simultaneous mapping of thousands of connections without the need for painstaking manual reconstruction or slow imaging processes.
Pilot Success: Mapping the Pontocerebellar Circuit
To demonstrate the efficacy of Connectome-seq, Zhao’s team focused on the pontocerebellar circuit in the mouse brain. This circuit is a vital pathway that links the pons—a part of the brainstem—to the cerebellum. It plays a critical role in motor coordination, sensory perception, and certain cognitive functions.
The team successfully mapped more than 1,000 neurons within this circuit. The results were startling: the high-resolution data revealed previously unknown patterns of connectivity. Specifically, the researchers found direct links between certain cell types that were not previously known to connect in the adult brain. These findings suggest that even well-studied regions of the brain may contain "hidden" circuits that traditional mapping methods have overlooked.
The ability to discover new connections in a known circuit highlights the sensitivity of Connectome-seq. Because the method relies on molecular signatures rather than visual identification, it can detect subtle or rare connections that might be missed by an exhausted human researcher or a computer vision algorithm struggling with overlapping fibers in a 3D image.
Implications for Alzheimer’s and Neurodegenerative Research
The most profound impact of Connectome-seq may lie in the study of neurodegenerative diseases. Conditions like Alzheimer’s, Parkinson’s, and Amyotrophic Lateral Sclerosis (ALS) are increasingly understood as "disconnections syndromes." In the early stages of these diseases, the physical structure of the neurons may remain intact, but the synapses—the communication points—begin to fail or disappear.
"With sequencing-based approaches, the time and cost are greatly reduced, which really makes it possible to see differences in different brains," Zhao said. This scalability is crucial. Because Connectome-seq is relatively fast and inexpensive compared to electron microscopy, researchers can compare the brain maps of healthy individuals with those of patients at various stages of disease progression.
In Alzheimer’s research, for example, there is a desperate need to identify the "first domino" to fall in the cascade of cognitive decline. Current diagnostic tools often detect the disease only after significant damage has occurred, such as the buildup of amyloid plaques or the death of large numbers of neurons. Connectome-seq could allow scientists to pinpoint the exact "weak links" in neural circuits before symptoms appear.
"If we can catch where exactly the weak link is that kick-starts the whole catastrophic cascade in Alzheimer’s disease, can we specifically strengthen those connections so the disease slows or does not progress?" Zhao asked. This "circuit-guided" approach to therapy could lead to treatments that target specific vulnerable pathways rather than the entire brain, potentially reducing side effects and increasing efficacy.
Addressing the Data Challenge and Future Scalability
While Connectome-seq offers a faster path to mapping, it generates a staggering amount of data. Sequencing thousands of neurons and their millions of potential connections requires sophisticated computational tools to organize and interpret the results. However, the researchers are optimistic. Unlike image-based data, which is bulky and difficult to process, sequencing data is inherently digital and fits well into existing bioinformatics pipelines used in genomics.
The research team is already working on improvements to the platform. Their ultimate goal is to scale the technology to map the entire mouse brain—a feat that would provide a foundational reference for all of neuroscience. A complete mouse connectome would allow researchers to see how different systems—such as memory, emotion, and motor control—interact at a granular level.
The project has garnered significant support from the scientific community. Funding was provided by a Neuro-omics Initiative grant from the Wu Tsai Neurosciences Institute at Stanford University, as well as the Elsa U. Pardee Foundation and the Edward Mallinckrodt Jr. Foundation. This interdisciplinary support reflects the growing consensus that the future of neuroscience lies at the intersection of biology, engineering, and data science.
A New Era of "Circuit-Guided" Interventions
As the field of connectomics matures, the focus is shifting from simply "drawing the map" to "using the map." The development of Connectome-seq aligns with a broader trend toward precision medicine in neurology. Just as genomic sequencing has allowed doctors to tailor cancer treatments to a patient’s specific genetic profile, connectomic sequencing could allow for personalized interventions for psychiatric and neurological disorders.
For instance, in cases of treatment-resistant depression or schizophrenia, identifying specific "miswired" circuits could allow for more precise application of Deep Brain Stimulation (DBS) or Transcranial Magnetic Stimulation (TMS). By knowing exactly which connections are disrupted, clinicians can target their interventions with surgical precision.
Furthermore, the technology provides a platform for testing new drugs. Pharmaceutical companies could use Connectome-seq to observe how a candidate drug affects synaptic connectivity in real-time, providing a much more detailed picture of a drug’s impact than behavioral observations alone.
Conclusion: The Brain as a Central Processing Unit
The work of Zhao and his colleagues reinforces a computational view of the brain. By comparing the brain to a computer’s central processing unit (CPU), Zhao emphasizes that understanding function is impossible without understanding the underlying circuitry.
"If you don’t know how everything is wired together, you can’t understand its function, optimize it or fix it when something breaks," Zhao noted. For decades, the "black box" of the brain has hindered the development of effective cures for many of our most devastating diseases. Connectome-seq provides the flashlight needed to look inside that box.
As Connectome-seq continues to be refined and adopted by the global scientific community, it promises to turn the mystery of neural connectivity into a manageable, searchable database. While the road to a complete map of the human brain remains long, the ability to rapidly sequence the connections of the mouse brain marks the beginning of a new chapter in our quest to understand the biological basis of thought, memory, and disease. The "sequencing revolution" that transformed genetics in the 1990s and 2000s has now arrived for the connectome, and the implications for human health are profound.










Leave a Reply