A groundbreaking discovery by scientists at the California Institute of Technology (Caltech) has illuminated a critical vulnerability in bacterial defenses, revealing how viruses that prey on bacteria have independently evolved remarkably similar strategies to disable a key protein essential for cell wall construction. This finding, published in the February 26 issue of the prestigious journal Nature, not only deepens our understanding of viral-bacterial interactions but also points towards a promising new avenue for the development of urgently needed antibiotics. The research, spearheaded by graduate student Yancheng Evelyn Li under the guidance of Professor Bil Clemons, a leading figure in biochemistry at Caltech, highlights the protein MurJ as a prime target for future therapeutic interventions.
The Escalating Crisis of Antibiotic Resistance
The urgent need for novel antibiotics cannot be overstated. Bacteria, with their rapid evolutionary capacity, are proving increasingly adept at evading existing drugs. This adaptability has fueled a global public health crisis, where once-treatable infections are becoming life-threatening. Professor Clemons articulates this challenge starkly: "Evolution is powerful, and in bacteria, resistance to antibiotics develops quickly. This means that we now deal with bacteria that are resistant to all the medicines that we have." The statistics are sobering: in the United States alone, tens of thousands of individuals succumb annually to infections caused by antibiotic-resistant bacteria, a figure projected to rise alarmingly. This escalating threat compels researchers worldwide to seek entirely new strategies and to identify previously unrecognized bacterial weaknesses.
Targeting the Bacterial Fortress: The Cell Wall
A long-standing focus in the quest for new antibiotics has been the intricate machinery bacteria employ to build their cell walls. Specifically, the peptidoglycan biosynthesis pathway has garnered significant attention. Peptidoglycan, a rigid polymer, forms the structural backbone of the bacterial cell wall, providing essential protection against osmotic pressure and maintaining cellular integrity. Its critical role makes it an attractive target because it is a feature unique to bacteria, absent in human cells. As Professor Clemons observes, "Peptidoglycan is a unique feature of bacteria, and that makes it an attractive antibiotic target."
Several established antibiotics, such as penicillin and its derivatives like amoxicillin, exert their therapeutic effect by interfering with later stages of peptidoglycan synthesis. However, the relentless evolution of resistance necessitates a deeper dive into the fundamental processes of cell wall construction to uncover new, exploitable vulnerabilities.
The Vital Trio: MraY, MurG, and MurJ
At the heart of peptidoglycan biosynthesis lies a crucial set of proteins responsible for transporting the essential building blocks across the bacterial inner membrane. Among these, MraY, MurG, and MurJ play pivotal roles. These proteins act as molecular escorts, shuttling the precursors needed for cell wall assembly from the cytoplasm to the exterior of the inner membrane. The failure of any one of these proteins would halt peptidoglycan production, ultimately leading to bacterial death. Consequently, they represent highly promising targets for drug development.
While considerable progress has been made in understanding the general functions of these proteins, specific mechanistic details have remained elusive. Professor Clemons acknowledges that "important mechanistic details remain unclear." Currently, no approved drugs directly inhibit MraY, MurG, or MurJ. Nevertheless, the potential for therapeutic intervention is significant. "We do know that we can find small molecules, either derived from nature or synthesized in chemical libraries, that will inhibit these proteins," Professor Clemons notes. Furthermore, recent discoveries have revealed that bacteriophages, viruses that infect bacteria, have independently evolved sophisticated mechanisms to target this very pathway.
Bacteriophages: Masters of Bacterial Invasion
Bacteriophages, or phages, are nature’s own bacterial adversaries. To propagate, they must successfully infect a bacterial cell, replicate within it, and then exit to infect new hosts. A critical step in this life cycle involves breaching the bacterial cell wall. Professor Clemons explains, "Getting back out means that they have to get past the peptidoglycan layer. Because it acts like chainmail, the phages get stuck if they can’t break through it." This inherent need to navigate the peptidoglycan layer makes phages adept at exploiting its vulnerabilities.
The Clemons laboratory focuses its research on small phages that possess compact genomes, often containing single-stranded DNA or RNA. These viruses employ streamlined strategies to overcome bacterial defenses. In a significant prior publication in Science in 2023, the team detailed their work on the well-studied phage ΦX174, a virus with a long history of research at Caltech.
Viral Proteins That Subdue MurJ
The strategy employed by these small phages to dismantle bacterial defenses often involves specialized proteins known as single-gene lysis proteins, or Sgls (pronounced "sigils"). These Sgls are encoded within the phage genome and are instrumental in the phage’s ability to lyse, or break open, the bacterial host. Li and Clemons have specifically investigated Sgls that target MurJ, one of the aforementioned key cell wall proteins.
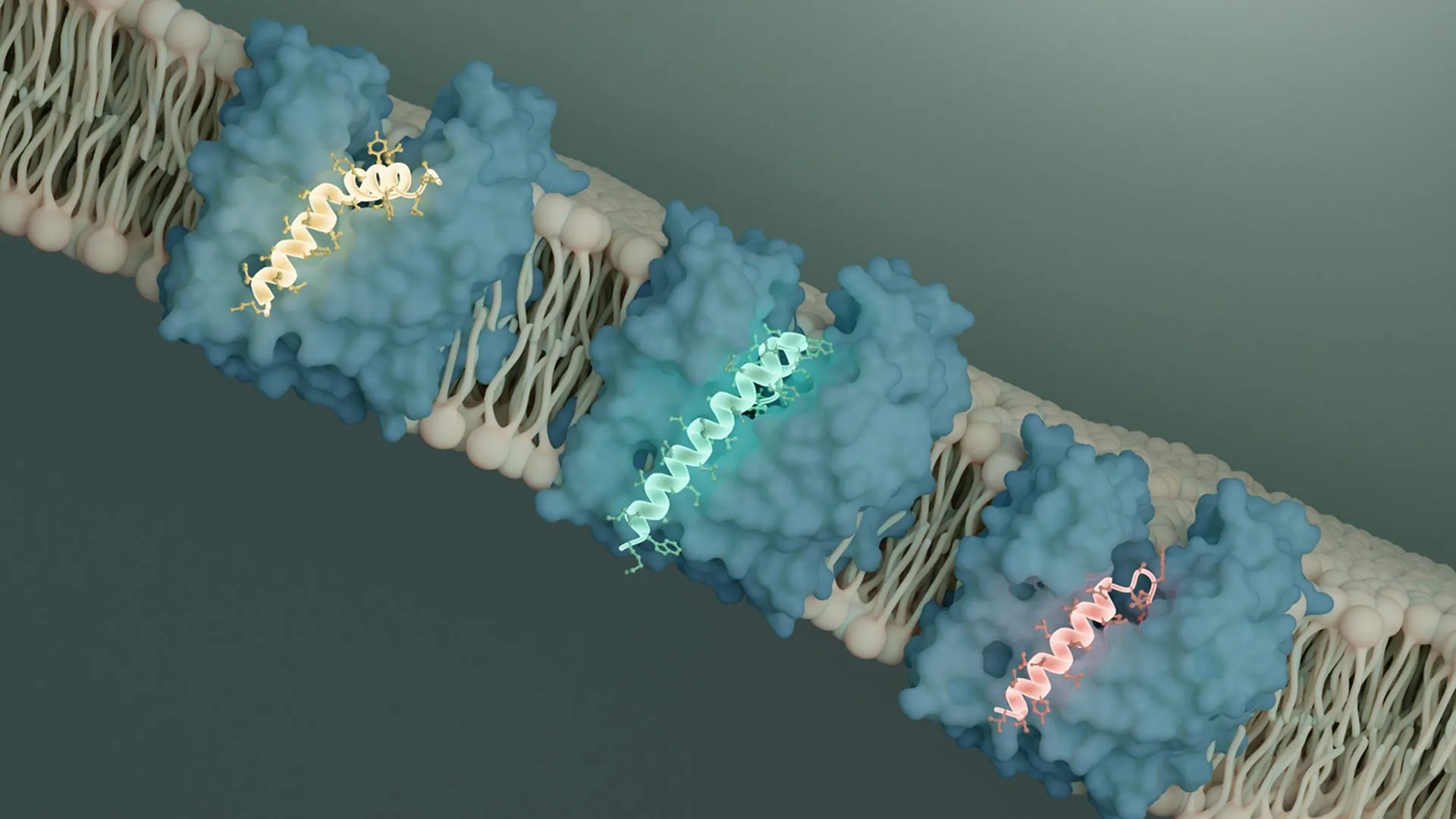
MurJ functions as a flippase, a critical enzyme that facilitates the translocation of peptidoglycan precursors across the bacterial inner membrane. It operates by undergoing conformational changes, alternately exposing the precursor to the inside and then the outside of the cell, without creating a permanent pore. This dynamic process ensures the efficient delivery of building materials for the cell wall. Earlier research by collaborators had demonstrated that two distinct and evolutionarily unrelated Sgls, known as SglM and SglPP7, both achieve bacterial lysis by inhibiting MurJ.
Unraveling the Mechanism of Inhibition
To elucidate how these viral proteins exert their inhibitory effect, Li employed state-of-the-art cryo-electron microscopy (cryo-EM) at Caltech’s Beckman Institute Biological and Cryogenic Transmission Electron Microscopy (Cryo-EM) Resource Center. This advanced technique allowed researchers to visualize the intricate molecular interactions between the Sgl proteins and MurJ.
Li’s findings revealed a striking convergence in the inhibitory mechanisms of SglM and SglPP7. Both viral proteins were observed to bind to a specific groove on the MurJ protein. This binding event effectively prevents MurJ from undergoing the necessary structural transformations required for its transport function.
"It is clear that both of these Sgls bind to MurJ in an outward-facing conformation, locking it into this position," Li stated. This observation is particularly encouraging for therapeutic development. The outward-facing conformation of MurJ is, by definition, more accessible to molecules in the extracellular environment, suggesting that drugs designed to target this specific state could be more readily delivered and effective.
Convergent Evolution: A Testament to MurJ’s Importance
The discovery that two unrelated phages have independently evolved to inhibit MurJ in such a similar manner was a significant surprise to the research team. "These peptides, which have no evolutionary links to each other, have both figured out how to target MurJ in a very similar way. These are two examples of convergent evolution, in which different evolutionary paths arrive at the same solution. We were surprised!" Professor Clemons remarked.
Convergent evolution, where distinct lineages independently evolve similar traits or solutions to similar problems, underscores the fundamental importance of MurJ as a bacterial target. The rapid evolutionary pace of viruses suggests that many more phages likely harbor similar Sgls capable of inhibiting MurJ. The ease with which phages can be isolated and their genomes sequenced presents a rich source for identifying novel biological insights and potential antibiotic targets.
A Third Viral Ally Reinforces the Target
Building on these compelling findings, the Caltech team, in collaboration with researchers at Texas A&M University, delved deeper into the phage genome landscape. In their Nature publication, they analyzed another phage genome and identified a novel Sgl, designated SglCJ3, originating from a phage predicted to be named Changjiang3. Using cryo-EM, Li again determined the structural complex of SglCJ3 bound to MurJ. The results were consistent with previous observations: SglCJ3 also locks MurJ in the same outward-facing conformation, effectively halting its function.
"This is a third genome that evolved a distinct peptide to inhibit the same target in a similar way," Professor Clemons emphasized. This repeated evolutionary solution provides "the first strong evidence that evolution identifies MurJ as a great target for killing bacteria." The implication is clear: researchers should "follow evolution’s lead and develop therapeutics that target MurJ." This discovery powerfully illustrates how fundamental biological research can pave the way for critical medical advancements. The team’s future path is firmly set on leveraging the insights gained from Sgl discovery, with a hopeful outlook for continued support to translate these foundational concepts into tangible therapeutic realities.
Broader Implications and Future Directions
The identification of MurJ as a conserved and evolutionarily validated target for bacterial disruption holds significant promise for the future of infectious disease treatment. The rapid emergence of antibiotic-resistant pathogens necessitates a paradigm shift in drug discovery, moving beyond modifications of existing drug classes to the identification of entirely new targets and mechanisms of action.
The study’s findings suggest a potential for developing small molecule inhibitors or even phage-derived therapies that mimic the action of Sgls. By precisely targeting MurJ in its outward-facing conformation, such therapeutics could offer a novel and effective means of combating a wide spectrum of bacterial infections, including those currently resistant to conventional antibiotics.
Furthermore, the research highlights the immense untapped potential of bacteriophages as a source of novel antimicrobial agents and biological insights. The ongoing exploration of diverse phage populations could yield a wealth of new Sgls and other antibacterial compounds, accelerating the discovery pipeline for much-needed new antibiotics.
Authorship and Funding
The foundational research detailed in the Nature paper, titled "Convergent MurJ Flippase Inhibition by Phage Lysis Proteins," was a collaborative effort involving key contributors from Caltech and Texas A&M University. In addition to lead authors Yancheng Evelyn Li and Bil Clemons, the author list includes Grace F. Baron, a graduate student at Caltech, and Francesca S. Antillon, Karthik Chamakura, and Ry Young from Texas A&M University.
This significant scientific endeavor was made possible through generous funding from several prominent organizations. Support was provided by the Chan Zuckerberg Initiative, the National Institutes of Health, the G. Harold and Leila Y. Mathers Foundation, and the Center for Phage Technology at Texas A&M, which is jointly sponsored by Texas A&M AgriLife. The multi-faceted support underscores the recognized importance and potential impact of this research in addressing the critical global challenge of antibiotic resistance.









Leave a Reply