A pioneering scientific breakthrough has emerged from Scripps Research in California, USA, in collaboration with IAVI (New York, USA) and other esteemed institutes: the development of a novel nanodisc platform that offers an unprecedentedly clear view of how crucial viral proteins engage with antibodies. This advancement, detailed in a recent publication in Nature Communications on February 10, 2026, marks a significant leap forward in vaccine research, particularly for notoriously challenging pathogens such as HIV and Ebola.
The Enduring Challenge of Viral Evasion and Vaccine Design
Viruses are intricate biological entities, adept at circumventing host immune defenses. A key component of their invasive strategy involves specialized proteins that adorn their surfaces, acting as keys to unlock cellular entry. These surface proteins, often embedded within the viral membrane, are the primary targets for vaccine development. The traditional approach to vaccine design frequently involves synthesizing laboratory-made versions of these viral surface proteins to elicit an immune response. However, a persistent challenge has plagued this methodology: these lab-produced proteins typically lack critical membrane-anchoring regions, meaning they do not perfectly replicate the native conformation and environment found on a real virus. This structural divergence has historically hindered a comprehensive understanding of how antibodies effectively identify, bind to, and neutralize these viral targets in their natural state.
For decades, vaccine scientists have grappled with this fidelity issue. The absence of the membrane context can alter the protein’s three-dimensional structure, expose non-native epitopes (the parts of an antigen that are recognized by the immune system), or obscure crucial epitopes that are only accessible when the protein is correctly oriented within a lipid bilayer. Such discrepancies complicate the design of vaccines capable of inducing broadly protective, long-lasting immunity. This is especially true for viruses like HIV, which exhibit high genetic variability and employ sophisticated mechanisms to evade antibody recognition, making the precise targeting of conserved, functionally critical regions paramount.
Introducing the Nanodisc Solution: A Native-Like Environment
Addressing this fundamental limitation, the team at Scripps Research and IAVI has engineered a platform that enables the study of viral surface proteins in a form that remarkably approximates their natural appearance on an intact virus. The core of this innovative approach lies in the utilization of nanodisc technology. These viral proteins are meticulously embedded into nanoscale particles composed of lipid molecules, thereby preserving them within a membrane-like structure. This artificial, yet highly realistic, environment allows researchers to observe the intricate dance between viral proteins and antibodies with unparalleled accuracy, providing crucial insights that could fundamentally reshape vaccine development strategies.
"For many years, we’ve had to rely on versions of viral proteins that are missing important pieces," explained co-senior author William Schief, a professor at Scripps Research and executive director of vaccine design at IAVI’s Neutralizing Antibody Center. "Our platform lets us study these proteins in a setting that better reflects their natural environment, which is critical if we want to understand how protective antibodies recognize a virus." This statement underscores the paradigm shift the nanodisc platform represents, moving from simplified models to physiologically relevant representations.
A Deeper Dive into Nanodisc Technology and Its Advantages
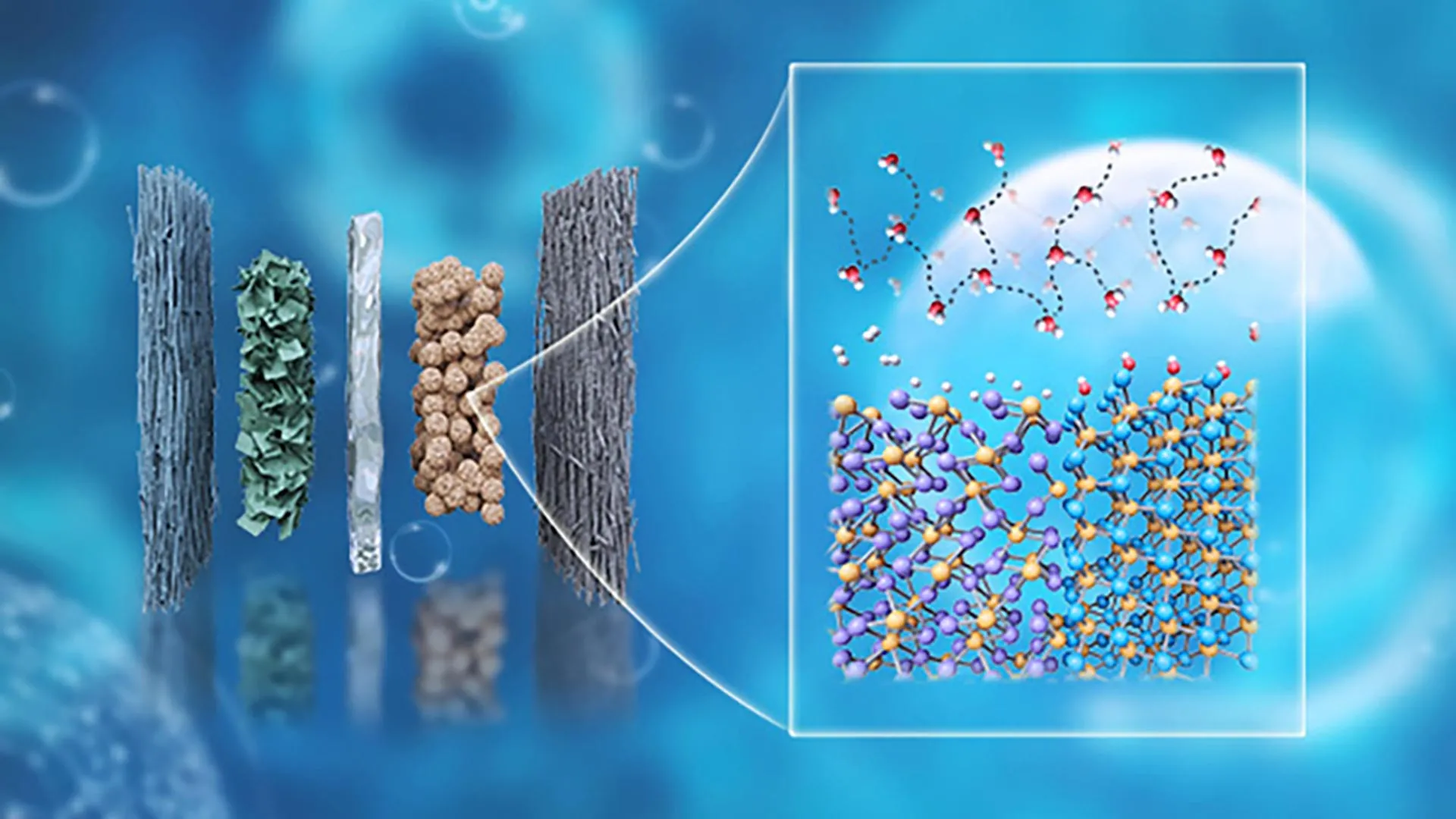
Nanodiscs are essentially miniature, discoidal patches of lipid bilayer, stabilized by membrane scaffold proteins (MSPs). These structures mimic cellular membranes, providing a stable, soluble, and native-like environment for integral membrane proteins. While the concept of nanodiscs has existed for some time, the innovation here lies in their specific application and optimization for complex viral surface proteins in the context of vaccine research. By encapsulating viral proteins within these lipid discs, the research team ensures that the proteins retain their native orientation, conformation, and accessibility, particularly at the membrane-proximal regions often crucial for broad antibody recognition.
In real viruses, surface proteins are not merely free-floating entities; they are intricately anchored within a lipid membrane and organized into specific tertiary and quaternary structures. This membrane embedding is vital for their function and for how they are presented to the immune system. Previous lab studies often necessitated the removal of the membrane-anchoring region to facilitate protein production and analysis, a pragmatic compromise that, while useful, inadvertently masked critical features, particularly for antibodies targeting regions near the base of the protein, adjacent to the viral membrane. The nanodisc platform effectively overcomes this compromise, offering a ‘true-to-life’ stage for these molecular interactions.
"Putting all of these components together into a single, reliable system was the key," added first author Kimmo Rantalainen, a senior scientist in Schief’s lab. "The individual pieces already existed, but making them work together in a way that’s reproducible and scalable opens up new possibilities for how vaccines are analyzed and designed." This highlights the engineering prowess required to integrate existing nanodisc technology with advanced viral protein expression and immunological assay techniques into a cohesive, functional pipeline.
Pioneering Applications: Tackling HIV and Ebola

To validate the robustness and efficacy of their novel platform, the researchers applied it to proteins from HIV and Ebola. These two viruses represent formidable challenges for vaccine developers due to their complex surface glycoproteins, high mutation rates, and the difficulty in eliciting broadly neutralizing antibodies (bNAbs).
For HIV, the team specifically focused on a conserved region of the virus’s envelope glycoprotein (Env) that sits near the membrane. This region is a known target for a class of bNAbs capable of neutralizing a broad spectrum of HIV variants. Such antibodies are highly sought after in HIV vaccine research because they recognize viral parts that remain stable even as the virus mutates, offering the promise of durable, pan-variant protection – an immune response that scientists ardently hope future vaccines can trigger.
The nanodisc platform proved instrumental in capturing detailed structural snapshots of how these crucial antibodies interact with the HIV Env protein in its native membrane context. These investigations revealed previously invisible features and interactions at the membrane interface, providing critical insights into antibody function that were unattainable when the protein was studied in isolation. These revelations are not merely academic; they offer mechanistic explanations for how certain antibodies neutralize the virus, often by destabilizing the protein structures it employs to infect cells. This knowledge is invaluable for guiding the rational design of future vaccines aimed at eliciting similar potent immune responses.
"The structure gave us a level of detail we simply couldn’t access before," noted Rantalainen. "It showed us new interactions at the membrane interface and suggested why those matter for antibody function." This level of atomic detail is crucial for structure-guided vaccine design, allowing researchers to precisely engineer immunogens that present desired epitopes to the immune system.
Beyond HIV, the team further demonstrated the broad applicability of their nanodisc platform by successfully applying it to Ebola proteins. This confirmed that antibodies could indeed identify and bind to these proteins within the same membrane-like environment, underscoring the versatility of the technology for a range of membrane-enveloped viruses.
Accelerating Vaccine Research and Development
The utility of this nanodisc platform extends far beyond basic structural studies. It is poised to revolutionize the analysis of immune responses to vaccine candidates. By employing the nanodiscs as molecular ‘bait,’ researchers can efficiently isolate and study specific immune cells, such as B cells, that recognize viral proteins. This provides a significantly clearer and more accurate picture of how the body responds to a given vaccine candidate, allowing for more informed decisions in preclinical and clinical development.
A critical advantage highlighted by the research team is the platform’s scalability and efficiency. What once required a month or even longer to prepare and analyze using conventional methods can now be accomplished in approximately a week. This dramatic reduction in preparation time makes the platform eminently practical for high-throughput screening and for systematically comparing multiple vaccine candidate designs side-by-side. This accelerated pace is vital in the fast-evolving landscape of infectious disease research and pandemic preparedness.
While the nanodisc platform itself is not a vaccine, its role as a sophisticated tool is undeniable. It promises to inform and significantly accelerate vaccine research, particularly for those viruses where traditional approaches have repeatedly fallen short. The ability to accurately mimic the viral surface in an ex vivo setting allows for early-stage validation of vaccine concepts, potentially reducing the high attrition rate of candidates in later clinical trials.
Broader Implications for Global Health and Future Outlook
The implications of this breakthrough stretch far beyond HIV and Ebola. The approach is broadly applicable to other viruses possessing similar membrane-embedded proteins, including globally prevalent pathogens like influenza and SARS-CoV-2. For influenza, which constantly mutates, and for emerging coronaviruses, the ability to present native-like antigens could lead to the development of more universal or pan-variant vaccines, offering broader and more durable protection.
This technology offers a more realistic and accurate means to test scientific hypotheses and vaccine concepts early in the development pipeline. By providing a faithful representation of viral surface proteins, it allows for a more precise identification of desirable antibody responses and a better understanding of how vaccine-induced antibodies neutralize pathogens. This could lead to:
- Enhanced Vaccine Efficacy: Designing vaccines that elicit antibodies targeting functionally critical and conserved epitopes, leading to more potent and long-lasting immunity.
- Faster Development Cycles: The increased efficiency and scalability mean vaccine candidates can be evaluated more quickly, shortening the timeline from discovery to clinical trials.
- Targeting Intractable Viruses: Offering new hope for diseases like HIV, for which a highly effective vaccine remains elusive despite decades of intensive research.
- Pandemic Preparedness: Equipping researchers with a powerful tool to rapidly develop and assess vaccine candidates against novel or emerging viral threats.
"This gives the field a more realistic, accurate way to test ideas early on," emphasized Schief. "By improving how we study viral proteins and antibody responses, we hope this platform will help advance next-generation vaccines against some of the world’s most challenging viruses." The collaborative nature of this research, bringing together expertise from Scripps Research and IAVI, highlights the power of inter-institutional partnerships in tackling complex biomedical challenges. As the scientific community continues to grapple with existing and emerging infectious diseases, tools like the nanodisc platform represent critical investments in global health security, paving the way for a new era of vaccine design.












Leave a Reply