Unforeseen Genetic Anomaly Discovered in Oxford Pond Organism
Oxford, UK – A routine scientific endeavor to test the capabilities of advanced single-cell DNA sequencing technology has yielded a discovery of profound significance, revealing a microscopic organism with a fundamentally altered genetic code. The organism, a previously unknown species of protist collected from a freshwater pond within Oxford University Parks, exhibits a unique interpretation of DNA’s fundamental instructions, a variation previously unseen by the scientific community.
The groundbreaking research, spearheaded by Dr. Jamie McGowan, a postdoctoral scientist at the Earlham Institute, aimed to rigorously evaluate a new DNA sequencing pipeline designed to function with exceptionally minute quantities of genetic material, including that extracted from a single cell. This practical objective, however, led the research team down an unexpected path, uncovering a biological marvel that challenges long-held assumptions about the universality of the genetic code.
A Serendipitous Discovery in the Microcosm
The genesis of this remarkable finding can be traced back to a seemingly ordinary research project. Dr. McGowan and his colleagues were focused on optimizing their DNA sequencing capabilities, a critical area for advancing fields such as personalized medicine and understanding microbial diversity. The choice of a protist from Oxford University Parks for this validation was, as Dr. McGowan himself noted, "sheer luck." The organism, now cataloged as Oligohymenophorea sp. PL0344, was not the primary subject of their investigation but rather a test case for their technological prowess.
Protists, a diverse group of eukaryotic microorganisms, are notoriously challenging to categorize due to their immense variability. They encompass a wide spectrum of life forms, from single-celled amoebas and algae to more complex multicellular organisms like kelp. "The definition of a protist is loose," Dr. McGowan explained in an interview, "essentially it is any eukaryotic organism which is not an animal, plant, or fungus. This is obviously very general, and that’s because protists are an extremely variable group. Some are more closely related to animals, some more closely related to plants." This inherent variability, while making them difficult to define, also makes them fertile ground for evolutionary innovation.
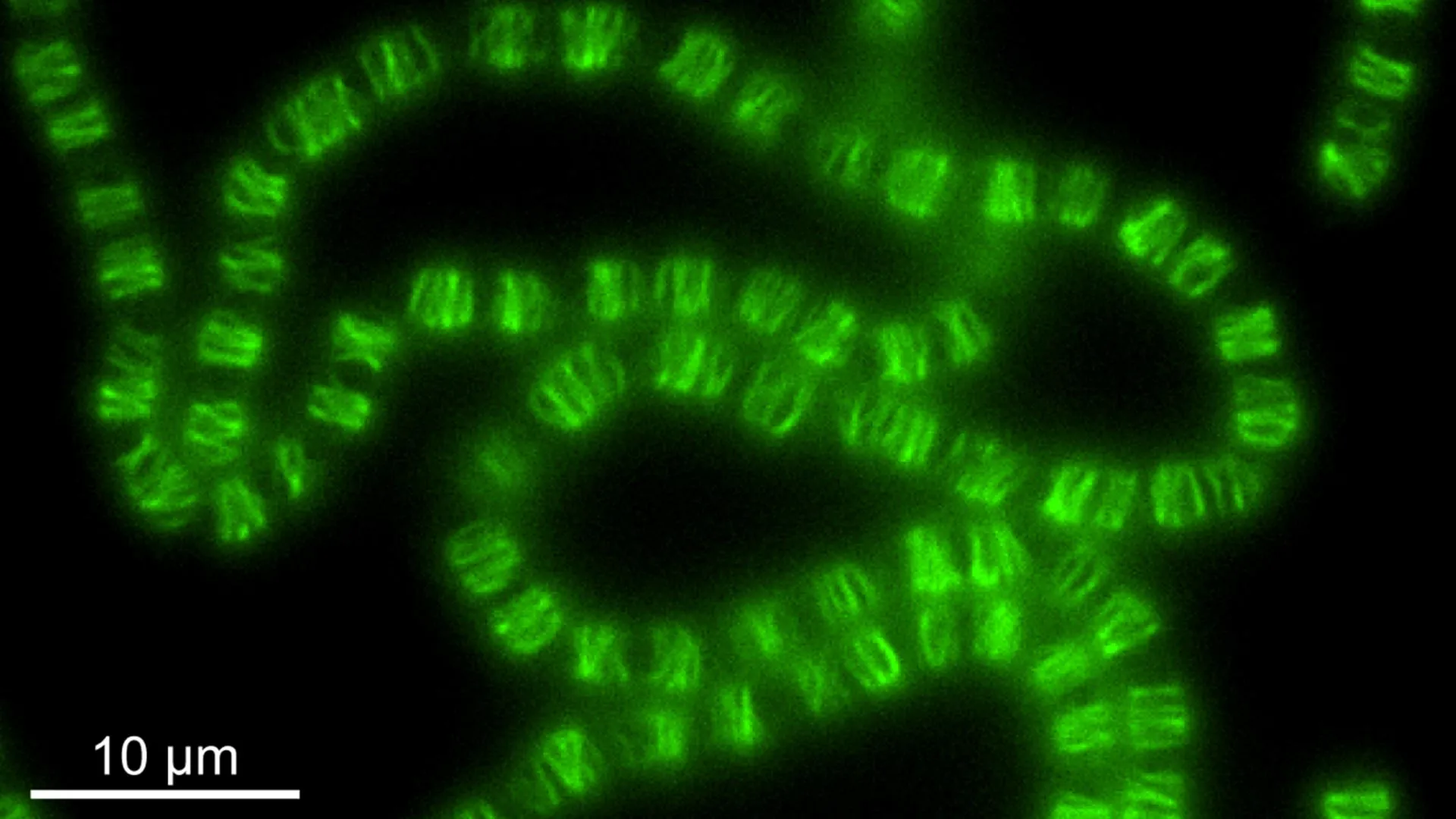
Oligohymenophorea sp. PL0344 belongs to a specific group of protists known as ciliates. These microscopic, single-celled organisms are characterized by their hair-like appendages called cilia, which they use for locomotion and feeding. Ciliates are ubiquitous in aquatic environments and have long been of interest to geneticists due to their propensity for genetic code alterations, particularly involving the "stop codons" that signal the termination of protein synthesis.
Reinterpreting the Language of Life: The Genetic Code Anomaly
The core of the discovery lies in the organism’s unusual interpretation of genetic instructions. In virtually all known life forms, the process of building proteins from DNA involves a set of three specific sequences, known as stop codons: TAA, TAG, and TGA. These triplets act as punctuation marks, signaling to the cellular machinery when to cease the assembly of amino acids, thereby completing a functional protein. The genetic code is widely regarded as "nearly universal" due to the consistent interpretation of these codons across the vast majority of species.
However, Oligohymenophorea sp. PL0344 has redefined this universality. The research, published in the esteemed journal PLOS Genetics, revealed that this specific protist has rewired its genetic language. Instead of all three acting as stop signals, two of these codons, TAA and TAG, have been reassigned to specify amino acids. Specifically, TAA now codes for lysine, and TAG directs the incorporation of glutamic acid. This is a departure from the established pattern where any known variation involving TAA and TAG typically sees them change in tandem and assign the same amino acid, suggesting an ancient evolutionary link.
"In almost every other case we know of, TAA and TAG change in tandem," Dr. McGowan elaborated. "When they aren’t stop codons, they each specify the same amino acid. This is extremely unusual. We’re not aware of any other case where these stop codons are linked to two different amino acids. It breaks some of the rules we thought we knew about gene translation – these two codons were thought to be coupled."
The research team also observed a higher-than-usual abundance of the TGA codon, which appears to have retained its function as a stop signal. This increased reliance on TGA may serve as a compensatory mechanism, ensuring the efficient termination of protein synthesis despite the altered roles of TAA and TAG. The study noted that the remaining UGA stop codon is frequently found positioned immediately after coding regions, suggesting a role in preventing "readthrough" – a phenomenon where translation continues beyond the intended stop signal, potentially leading to non-functional proteins.
The Mechanism of Translation and the Role of tRNA
The intricate process of converting genetic information into functional proteins is a cornerstone of molecular biology. DNA, the blueprint of life, is first transcribed into messenger RNA (mRNA). This mRNA molecule then travels to ribosomes, the cell’s protein-making factories, where it undergoes translation. During translation, the ribosome reads the mRNA sequence in three-nucleotide units called codons. Each codon specifies a particular amino acid, or in the case of stop codons, signals the end of the protein chain.
The discovery in Oligohymenophorea sp. PL0344 demonstrates that the cellular machinery, particularly transfer RNA (tRNA) molecules, has adapted to recognize and interpret these reassigned codons. The study’s genome and transcriptome analysis identified specific suppressor tRNA genes that precisely match the newly assigned codons, providing robust evidence that the organism genuinely reads TAA as lysine and TAG as glutamic acid. This finding underscores the plasticity of the cellular translation apparatus and its capacity to evolve novel interpretations of the genetic lexicon.
A Pattern of Genetic Innovation in Ciliates
This remarkable discovery is not an isolated incident. Subsequent research has further illuminated the tendency of ciliates to exhibit unusual genetic code variations. A 2024 study, also published in PLOS Genetics, reported multiple independent instances of the UAG (the RNA equivalent of TAG) stop codon being reassigned in other groups of ciliates, known as phyllopharyngeans. In some of these uncultivated ciliates, UAG was found to encode leucine, while in others, it specified glutamine.
Notably, in these phyllopharyngean ciliates, the UAA codon remained the preferred stop codon, mirroring the situation in Oligohymenophorea sp. PL0344. This pattern suggests that the reassignment of UAG into a protein-coding role has occurred repeatedly throughout ciliate evolution, reinforcing the notion that these microorganisms are significant exceptions to the standard genetic code. These findings collectively point towards a dynamic and adaptable genetic code, particularly within poorly understood microbial eukaryotes like ciliates, where evolution has actively "edited the instructions" in remarkable ways.
Broader Implications for Evolutionary Biology and Biotechnology
The implications of this discovery extend far beyond the confines of a single pond organism. It fundamentally challenges the long-held belief in the near-absolute universality of the genetic code, a concept that has been a bedrock of molecular biology for decades. The existence of such significant variations in natural systems opens new avenues for understanding the evolutionary history of life and the mechanisms by which genetic codes can change.
"Scientists attempt to engineer new genetic codes – but they are also out there in nature," Dr. McGowan remarked. "There are fascinating things we can find, if we look for them. Or, in this case, when we are not looking for them." This sentiment highlights the serendipitous nature of scientific discovery and the vast unknown that still exists within the microbial world.
From a biotechnology perspective, understanding how natural systems have evolved to manipulate the genetic code could have profound implications. It could inform efforts to engineer synthetic genetic codes for novel biotechnological applications, such as creating entirely new proteins or developing organisms with enhanced capabilities. The ability to precisely control and modify the genetic language, as demonstrated by Oligohymenophorea sp. PL0344, represents a frontier in synthetic biology.
A Timeline of Discovery
- Early 2023: Dr. Jamie McGowan and his team at the Earlham Institute initiate a project to test a new single-cell DNA sequencing pipeline.
- Mid-2023: During the testing phase, an unusual genetic anomaly is detected in a protist sample collected from Oxford University Parks.
- Late 2023: Rigorous analysis confirms that the protist, identified as Oligohymenophorea sp. PL0344, possesses a novel interpretation of genetic stop codons.
- December 2023: The findings are published in the peer-reviewed journal PLOS Genetics, detailing the reassignment of two stop codons.
- Early 2024: A subsequent study, also in PLOS Genetics, further strengthens the hypothesis that ciliates are frequent sites of genetic code variation, reporting multiple independent reassignment events in other ciliate species.
Funding and Future Research
The original research leading to this discovery was supported by significant funding, including the Wellcome Trust as part of the Darwin Tree of Life Project, and the Earlham Institute’s core funding from the Biotechnology and Biological Sciences Research Council (BBSRC), a part of UKRI. The genomic and transcriptomic data generated from this study have been made publicly available in established repositories, facilitating further research by the global scientific community.
The discovery of Oligohymenophorea sp. PL0344’s unique genetic code serves as a potent reminder of the unexplored biodiversity and biochemical novelty that lies hidden within our planet’s ecosystems. Future research will likely focus on elucidating the evolutionary pressures that led to these genetic code alterations and exploring the full extent of genetic code plasticity across the tree of life. The findings underscore the critical importance of continued exploration and technological advancement in unlocking the secrets of the microbial world.








Leave a Reply