A groundbreaking discovery by a research team led by Universitat Autònoma de Barcelona (UAB) scientist David Reverter has illuminated the intricate molecular machinery that orchestrates bacterial cell division. Published in the prestigious journal Nature Communications, the study precisely identifies how the MraZ protein interacts with the critical dcw gene cluster, revealing a fundamental regulatory mechanism essential for bacterial proliferation. This breakthrough provides unprecedented insight into a process vital for life, with far-reaching implications for understanding bacterial growth, development, and the potential for therapeutic interventions.
The Essential Dance of Cell Division
Cell division, the fundamental process by which a single cell gives rise to two or more daughter cells, is the cornerstone of life for all organisms, from the simplest bacteria to complex multicellular beings. This seemingly straightforward act is, in reality, a highly coordinated and tightly regulated symphony of molecular events. In the realm of bacteria, this intricate dance is largely choreographed by a conserved set of genes known as the dcw operon. This genetic ensemble serves as the blueprint, encoding the production of essential proteins that not only facilitate the physical act of cell division but also are instrumental in constructing the rigid scaffolding of the bacterial cell wall, the protective outer layer that defines bacterial shape and integrity.
The activation of these crucial genes within the dcw operon is precisely controlled by specialized proteins called transcription factors. These molecular gatekeepers bind to specific DNA sequences known as promoters, which act as the starting line for gene expression. The promoter region, situated just upstream of the gene’s coding sequence, signals where the cellular machinery should begin transcribing the genetic information into messenger RNA, the precursor to protein synthesis.
MraZ: The Master Conductor of Bacterial Replication
Central to this regulatory network is the MraZ protein. Remarkably, MraZ is itself the very first gene encoded within the dcw operon across virtually all bacterial species. This strategic placement underscores its pivotal role as the primary regulator. When MraZ is activated, it initiates a cascade of gene expression, triggering the production of the suite of proteins indispensable for bacteria to divide and multiply. In essence, MraZ acts as the master conductor, dictating the tempo and execution of the cell division process for the vast majority of bacterial populations. Understanding how MraZ exerts this control has been a long-standing quest in microbiology.
Illuminating the Molecular Mechanism: A Glimpse at Atomic Resolution
The UAB research group, under the leadership of Professor David Reverter of the Department of Biochemistry and Molecular Biology and a key researcher at the Institute of Biotechnology and Biomedicine of the UAB (IBB-UAB), has now provided an unprecedentedly detailed view of this regulatory mechanism. Employing state-of-the-art structural biology techniques, including X-ray crystallography and cryo-electron microscopy (cryo-EM), the scientists were able to visualize the molecular interactions with remarkable clarity.
These advanced imaging modalities allowed the team to meticulously map how the MraZ transcription factor engages with the promoter region of the dcw operon in Mycoplasma genitalium. This particular bacterium was chosen for its exceptionally compact genome, making it an ideal model organism for detailed genetic and molecular studies. The simplicity of its genetic makeup allows researchers to isolate and study specific pathways with greater precision.
The Dynamic Deformation of MraZ: A Surprising Revelation
The promoter region of the dcw operon, the specific DNA sequence targeted by MraZ, is characterized by the presence of four repetitive segments, often referred to as "boxes." Each of these boxes is composed of a specific sequence of six nucleotides, forming a critical recognition site for regulatory proteins. These repeated DNA sequences are fundamental to the precise control of gene transcription.
Through the power of cryo-electron microscopy, the researchers achieved nearly atomic resolution in observing the intricate interplay between the MraZ protein and the DNA bases within these four repetitive boxes. Their findings revealed a crucial and unexpected aspect of this interaction: MraZ must undergo a significant structural transformation, a dynamic deformation, to successfully bind to the promoter and initiate gene activation.
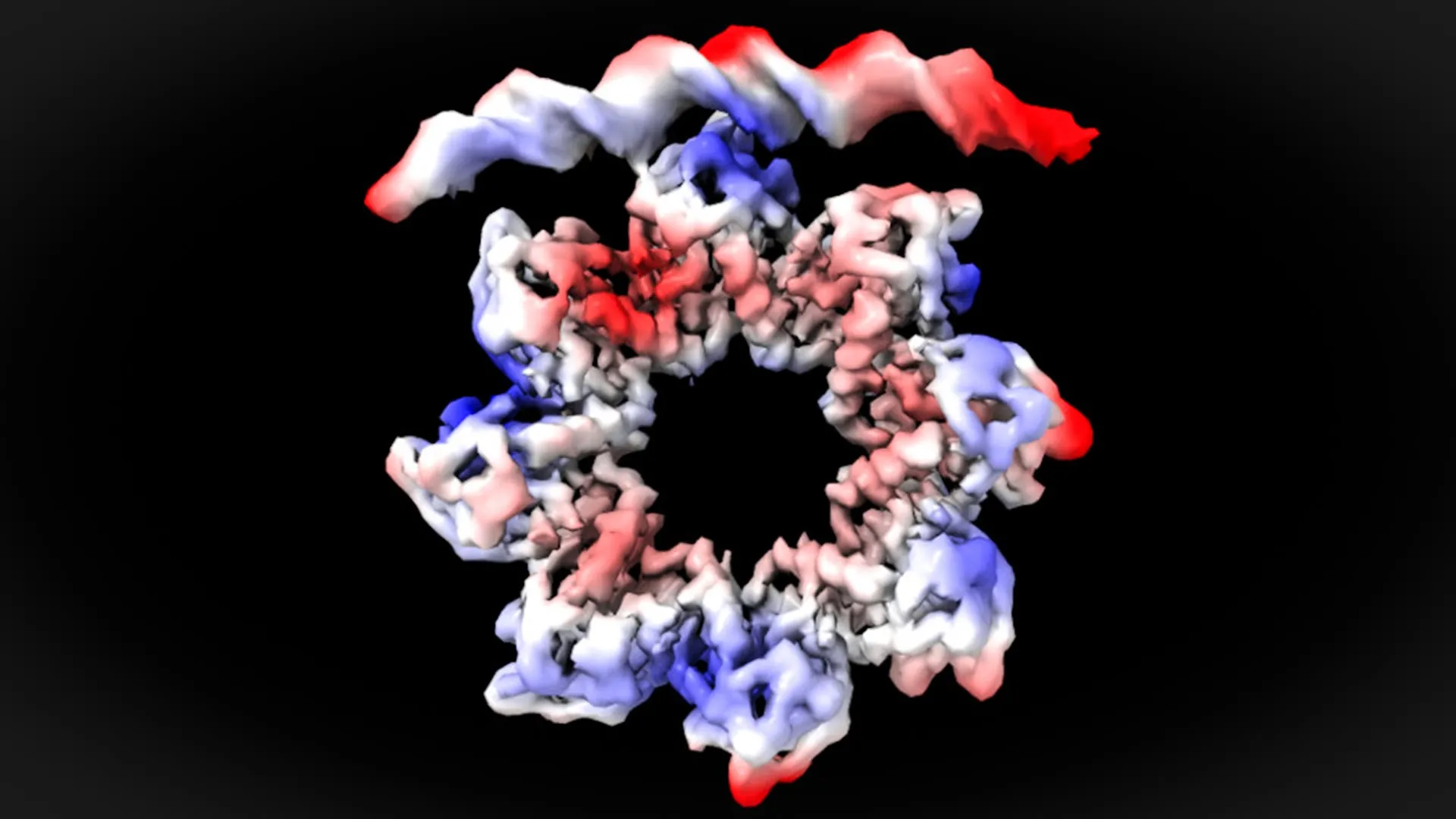
"This is a surprising observation," stated David Reverter. "The MraZ protein typically exists as an octamer, a complex formed by eight identical subunits arranged in a donut-like shape. However, this inherent structure, with its curvature, would seemingly preclude direct binding to the four distinct ‘boxes’ of the promoter. Yet, in order to regulate cell division, we observe a remarkable phenomenon: the donut ‘breaks’ and deforms in such a way that four of its subunits can precisely engage with the four DNA boxes of the promoter."
This dynamic reconfiguration of the MraZ octamer is a key finding, demonstrating that the protein is not a rigid entity but a flexible molecular machine capable of adapting its form to fulfill its regulatory function. This plasticity is crucial for its ability to recognize and bind to the specific DNA sequences that control the initiation of cell division.
A Paradigm Shift in Understanding Bacterial Control
The direct visualization of MraZ’s interaction with the promoter DNA represents a significant leap forward in the scientific community’s understanding of bacterial cell division. Prior to this study, researchers relied heavily on indirect methods, such as biochemical experiments and sophisticated computer modeling, to infer the working of this critical regulatory system. While these approaches provided valuable insights, they lacked the direct, visual confirmation that structural biology can offer. The atomic-level imaging provided by this research offers a level of detail and certainty that was previously unattainable.
The implications of this discovery extend beyond the specific bacterium studied. Professor Reverter expressed confidence in the broader applicability of their findings. "The regulatory system identified in this study is likely widespread among bacteria," he concluded. "This is because all MraZ proteins exhibit a high degree of similarity in their structure, maintaining the same octameric configuration. Furthermore, the DNA sequences of the promoters within the operons that regulate cell division are also remarkably conserved across different bacterial species." This universality suggests that the mechanism unveiled by the UAB team could be a common thread in bacterial proliferation across diverse microbial ecosystems.
A Timeline of Discovery and Collaboration
The journey to this significant revelation involved a concerted effort over several years, with key milestones in experimental design and data acquisition:
- Early Research & Hypothesis Formation: Initial studies in the field established the dcw operon’s importance and identified MraZ as a critical transcription factor. Biochemical data suggested a complex interaction but lacked definitive structural evidence.
- Technological Advancements: The development and refinement of cryo-electron microscopy and the increased accessibility of advanced X-ray crystallography facilities provided the necessary tools for high-resolution structural determination.
- Sample Preparation and Optimization (circa 2018-2020): The research team focused on optimizing the purification of MraZ and the preparation of Mycoplasma genitalium DNA for structural analysis. This stage involved numerous trials to achieve stable protein-DNA complexes suitable for imaging.
- Data Acquisition (circa 2020-2022): Experiments were conducted at national and international synchrotron facilities for X-ray crystallography and at specialized cryo-EM centers. This phase involved collecting vast amounts of imaging data.
- Structure Determination and Analysis (circa 2022-2023): Sophisticated computational methods were employed to reconstruct the 3D structures from the raw imaging data. This was followed by detailed analysis of protein-DNA interactions, conformational changes, and the identification of key amino acid residues involved in binding.
- Publication in Nature Communications (early 2024): Following rigorous peer review, the findings were published, making the detailed mechanism accessible to the global scientific community.
This ambitious research project was not undertaken in isolation. It was a testament to robust international collaboration. The primary research was spearheaded by David Reverter’s dedicated team at the Institute of Biotechnology and Biomedicine and the Department of Biochemistry and Molecular Biology at UAB. Crucial support and expertise were provided by the ALBA synchrotron, Spain’s national synchrotron light facility, which enabled high-resolution X-ray crystallography data collection. Furthermore, the cryo-electron microscopy services of the prestigious Institute of Genetics and Molecular and Cellular Biology of Strasbourg, France, were instrumental in achieving the atomic-level resolution of the MraZ-DNA complex.
Broader Implications and Future Directions
The implications of this discovery are multifaceted, spanning fundamental biology, medicine, and biotechnology.
- Fundamental Microbiology: This work provides a definitive molecular explanation for a fundamental biological process that has long been understood in principle but not in its detailed mechanistic execution. It enriches our understanding of the evolutionary conservation of regulatory pathways in bacteria.
- Antibiotic Development: Bacteria are a constant threat to human health, with antibiotic resistance emerging as a critical global challenge. The dcw operon and its regulatory machinery represent potential targets for novel antimicrobial agents. By disrupting the ability of MraZ to bind to the dcw promoter, it may be possible to inhibit bacterial growth and division without directly harming host cells. This research lays the groundwork for designing such targeted therapies. The identification of the precise binding interface and the dynamic conformational changes involved could inform the rational design of molecules that specifically interfere with this interaction.
- Biotechnology and Synthetic Biology: A precise understanding of bacterial cell division mechanisms can also be leveraged in synthetic biology for engineering bacteria for specific industrial or research purposes. For example, controlling the rate of bacterial growth is crucial in many biotechnological applications, from the production of biofuels to the synthesis of pharmaceuticals.
- Evolutionary Biology: The conserved nature of the MraZ protein and the dcw operon across diverse bacterial lineages offers valuable insights into bacterial evolution. Studying variations in these elements across different species could shed light on how bacteria have adapted and diversified over millennia.
The UAB team’s detailed structural insights into MraZ’s remarkable deformation and subsequent binding to the dcw promoter opens new avenues for scientific inquiry. Future research will likely focus on exploring the dynamic regulatory network in other bacterial species, investigating the role of accessory proteins in modulating MraZ activity, and ultimately translating these fundamental discoveries into tangible applications for combating bacterial infections and advancing biotechnological innovation. The clarity provided by this research marks a significant turning point in our ability to understand and potentially manipulate the very essence of bacterial life.








Leave a Reply