In a landmark advancement for the field of neuroscience, researchers have unveiled a high-speed, high-resolution method for mapping the intricate web of connections within the brain. By utilizing molecular "barcodes" made of RNA, a team led by the University of Illinois Urbana-Champaign has developed a platform called Connectome-seq, which allows for the simultaneous tracking of thousands of neural pathways at the level of individual synapses. This breakthrough, recently published in the journal Nature Methods, represents a fundamental shift in how scientists visualize the brain’s "wiring diagram," moving from labor-intensive visual imaging to high-throughput genetic sequencing.
The brain is often compared to the world’s most sophisticated computer, yet for decades, scientists have lacked a complete "schematic" of its internal circuitry. While the general regions of the brain responsible for memory, motor skills, and emotion are well-known, the specific cell-to-cell connections—the synapses—remain largely mysterious due to their staggering complexity and microscopic scale. Connectome-seq seeks to bridge this gap, offering a scalable solution that could transform our understanding of neurodegenerative diseases, such as Alzheimer’s and Parkinson’s, by identifying exactly where and how brain networks begin to fail.
The Challenge of Traditional Connectomics
To appreciate the significance of Connectome-seq, one must consider the historical context of brain mapping, a field known as connectomics. For over a century, the gold standard for understanding neural architecture was established by pioneers like Santiago Ramón y Cajal, who used silver staining and manual sketches to document the morphology of neurons. In the modern era, this evolved into electron microscopy (EM), a process that involves slicing brain tissue into sections thinner than a human hair, imaging them with high-powered microscopes, and painstakingly reconstructing the 3D pathways of axons and dendrites.
While electron microscopy provides unparalleled detail, it is prohibitively slow and expensive. Mapping the full brain of a fruit fly—an organism with roughly 100,000 neurons—took an international consortium years of effort and millions of dollars. Applying such methods to the mouse brain (70 million neurons) or the human brain (86 billion neurons) remains an almost insurmountable computational and logistical challenge.
Alternative methods, such as viral tracing and fluorescent labeling, have allowed researchers to see where neurons "project" or send their signals. However, these tools often lack "synaptic resolution." They can show that a neuron in Region A reaches toward Region B, but they cannot confirm with certainty which specific cell in Region B it has formed a functional connection with. This lack of precision has hindered efforts to understand how specific circuits process information and how these circuits are disrupted in disease states.
Turning Biology into a Sequencing Problem
The core innovation of Connectome-seq, developed by Boxuan Zhao, a professor of cell and developmental biology at the University of Illinois Urbana-Champaign, is the translation of a physical mapping problem into a digital data problem. By leveraging the power of next-generation sequencing (NGS), the team has bypassed the need for exhaustive visual reconstruction.
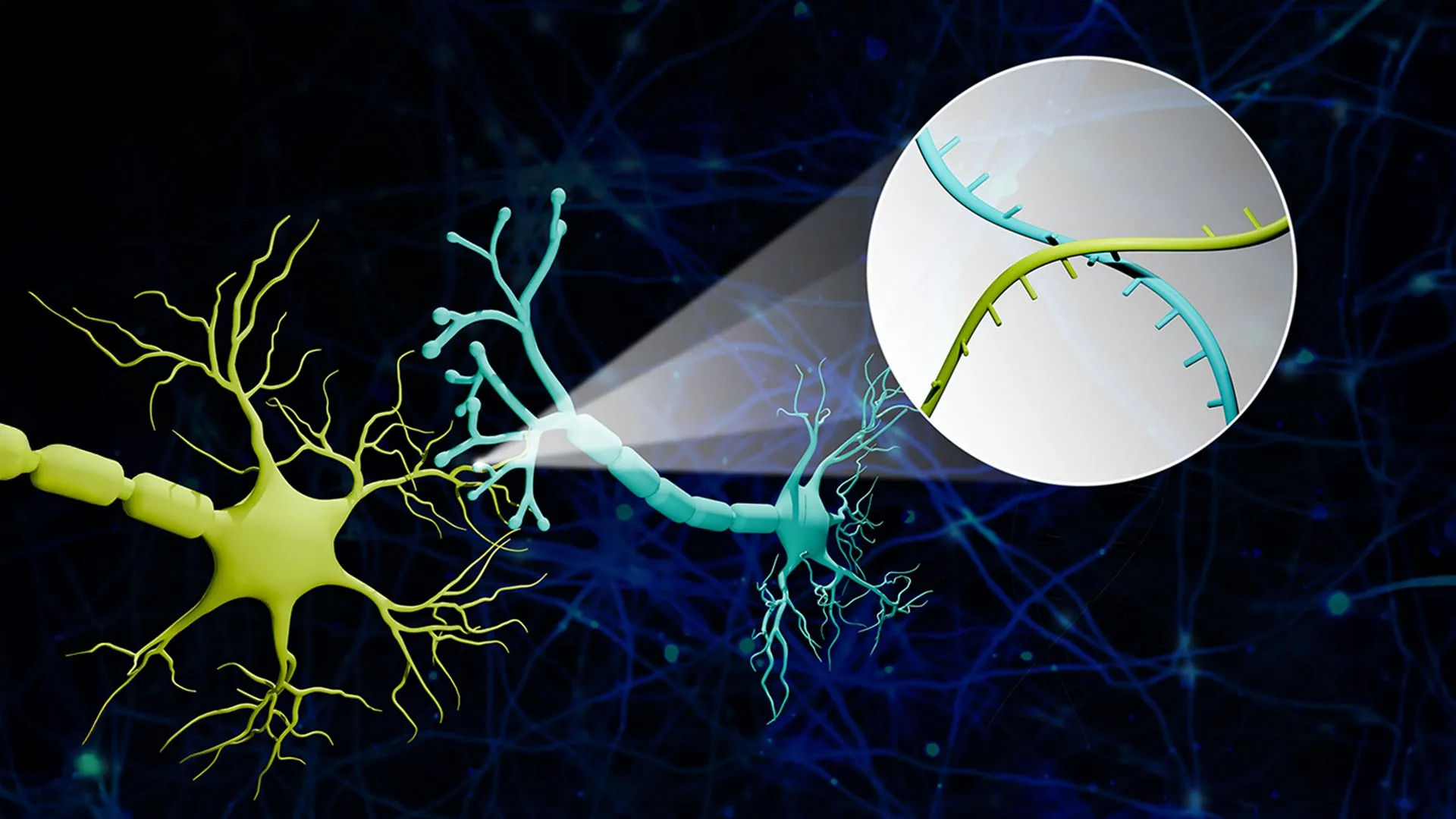
The process begins by assigning each individual neuron a unique molecular identity. Using engineered viral vectors, the researchers "tag" neurons with distinct RNA barcodes—essentially unique strings of genetic code that act as digital name tags. To solve the problem of identifying connections, the team engineered specialized proteins designed to transport these RNA barcodes from the cell body (the soma) down the long "cables" of the neuron (the axons) directly to the synapse.
"Imagine a big bunch of balloons," explained Zhao. "The main body of each balloon has its unique barcode stickers all over it, and some move down to the end of the string. If two balloons are tied together at the end, the two barcodes meet at the junction. Then we snip out the knots and sequence the barcodes in each one. If the same knot has stickers from balloon A and balloon B, we know these two balloons are tied together."
In the context of the brain, these "knots" are the synapses. By isolating the synaptic junctions and using high-throughput sequencing to read the barcode pairs found within them, the researchers can reconstruct a comprehensive map of which neurons are "talking" to each other. This method allows for the analysis of thousands of connections in a single experiment, a feat that would take months or years using traditional imaging techniques.
Experimental Validation: The Pontocerebellar Circuit
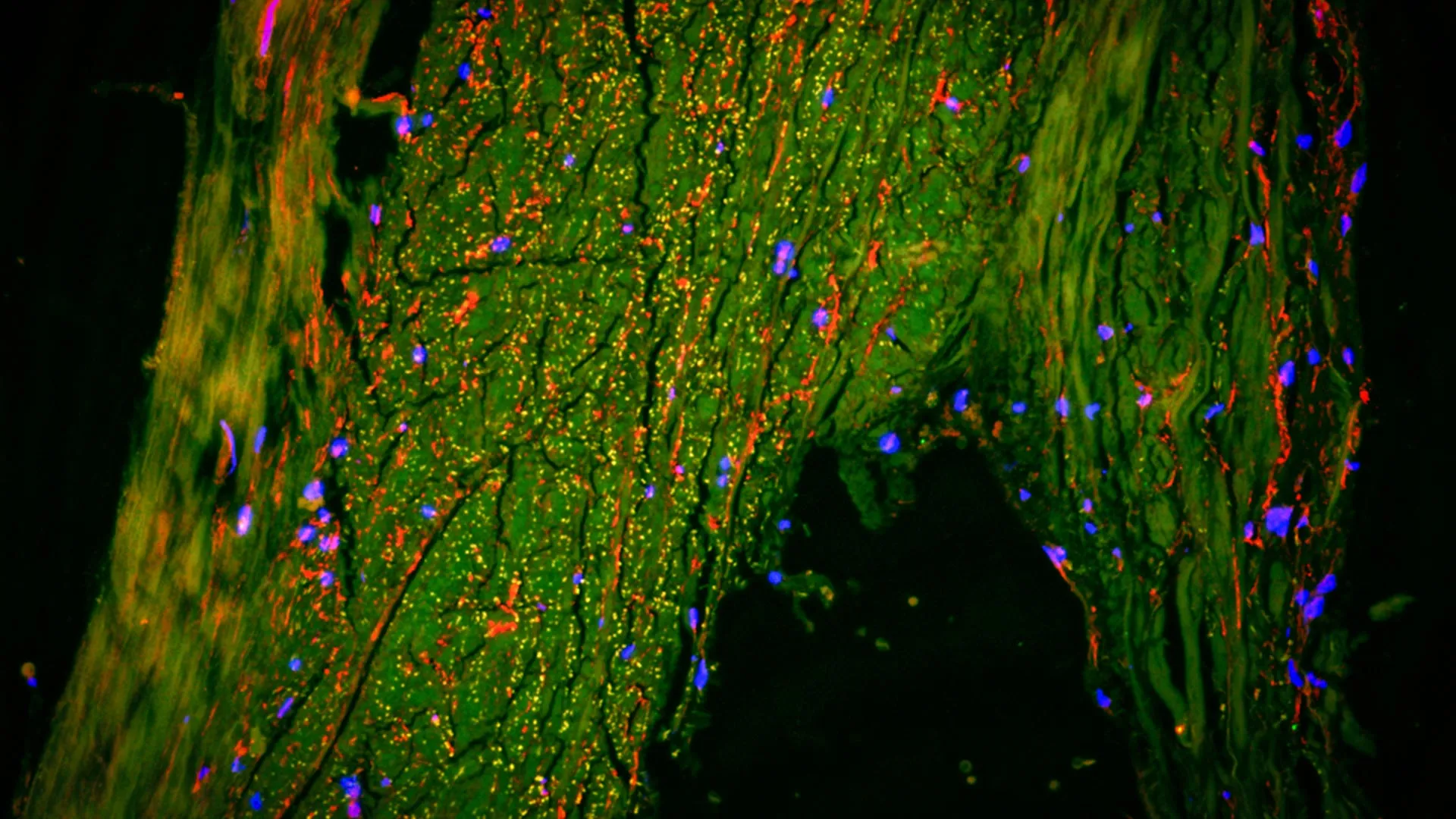
To demonstrate the efficacy of Connectome-seq, Zhao’s team applied the technology to the mouse brain, specifically targeting the pontocerebellar circuit. This pathway is a critical component of the motor system, linking the pons (a part of the brainstem) to the cerebellum, which coordinates movement and balance.
The results were both validating and surprising. The team successfully mapped over 1,000 neurons within this circuit, confirming established pathways while also discovering previously unknown connectivity patterns. Specifically, the analysis revealed direct links between certain cell types that had not been documented in the adult mouse brain. These findings suggest that the brain’s architecture may be even more complex and interconnected than current textbooks suggest.
The data gathered from the pontocerebellar experiment highlighted the "single-synapse resolution" of the platform. Unlike previous sequencing-based tools that provided a general "map of the neighborhood," Connectome-seq identifies the specific "house-to-house" connections. This level of detail is essential for understanding how the brain processes complex sensory input and executes precise motor commands.
Implications for Neurodegenerative Disease and Alzheimer’s Research
The most profound application of Connectome-seq lies in the study of brain disorders. Current research into Alzheimer’s disease, for instance, often focuses on the accumulation of amyloid plaques and tau tangles. However, many scientists believe that these are late-stage symptoms rather than the primary cause of cognitive decline. The "synaptoblastic" hypothesis suggests that the disease begins with the silent failure of specific neural circuits long before physical symptoms appear.
"With sequencing-based approaches, the time and cost are greatly reduced, which really makes it possible to see differences in different brains," Zhao noted. "We could see where connections change, where the most vulnerable parts of the brain are, perhaps before symptoms even appear."
If researchers can use Connectome-seq to compare the "wiring diagrams" of healthy brains against those in the early stages of Alzheimer’s, they may identify the "weak links" that trigger the catastrophic cascade of memory loss. This could pave the way for "circuit-guided therapeutic interventions"—treatments designed not just to clear plaques, but to specifically strengthen and repair the vital connections that maintain cognitive function.
Furthermore, the technology holds promise for psychiatric conditions such as schizophrenia and autism spectrum disorders, which are increasingly viewed as "connectopathies"—diseases characterized by atypical neural wiring rather than gross physical damage to the brain.
Scalability and the Future of the Whole-Brain Connectome
The research, which received support from the Wu Tsai Neurosciences Institute at Stanford University, the Elsa U. Pardee Foundation, and the Edward Mallinckrodt Jr. Foundation, is only the beginning. Zhao and his colleagues are already working on iterations of the platform to increase its throughput and sensitivity.
The ultimate goal of the lab is to map the entire mouse brain in its entirety. Achieving this would provide a baseline "atlas" for neuroscience, allowing researchers worldwide to test how various factors—ranging from environmental stress to new drug candidates—affect the brain’s global connectivity.
"With improvements already underway in our lab, we are confident that we can make it even better and eventually reach the goal of mapping the whole mouse brain," Zhao said.
The shift toward "neuro-omics"—the integration of genomics, proteomics, and connectomics—represents the next frontier of biological science. As the cost of genetic sequencing continues to drop, tools like Connectome-seq will become increasingly accessible to the broader scientific community. This democratization of brain mapping could lead to a surge in discoveries, as laboratories that previously lacked the resources for high-end electron microscopy can now utilize sequencing facilities to explore the neural circuits of various model organisms.
Analysis: A Paradigm Shift in Neuroscience
The development of Connectome-seq marks a departure from the "visual" era of neuroscience into the "data-driven" era. By treating the brain as a sequence-able entity, researchers have effectively applied the logic of the Human Genome Project to the structure of the mind.
One of the key advantages of this sequencing-based approach is its compatibility with other molecular tools. Because the "map" is made of RNA, it can theoretically be cross-referenced with "transcriptomic" data, which tells researchers which genes are active in a specific neuron. This means scientists could soon see not only which cells are connected, but also the functional state of those cells—for example, whether a specific connection is excitatory, inhibitory, or currently under stress.
However, challenges remain. While Connectome-seq is exceptionally fast at identifying connections, it does not yet provide the same level of spatial context as imaging. Researchers know which cells are connected, but they may lose some information about the physical shape and location of those connections within the brain’s 3D space. Future versions of the technology will likely seek to integrate sequencing data with spatial transcriptomics to create a truly multidimensional map.
Despite these hurdles, the consensus among the scientific community is one of cautious optimism. The ability to map thousands of connections simultaneously at single-synapse resolution is a capability that simply did not exist until now. As Boxuan Zhao emphasized, you cannot fix a broken computer if you don’t understand the circuitry. With Connectome-seq, humanity has taken a significant step toward finally understanding the circuitry of the most complex machine in the known universe.










Leave a Reply